[English] 日本語
 Yorodumi
Yorodumi- EMDB-24969: Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412) -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412) | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HCoV / spike / OC43 / polyclonal antibody / EMPEM / VIRAL PROTEIN / VIRAL PROTEIN-Immune System complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / membrane Similarity search - Function | |||||||||
| Biological species |  Human coronavirus OC43 / Human coronavirus OC43 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
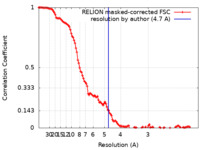
| Method | single particle reconstruction / cryo EM / Resolution: 4.7 Å | |||||||||
 Authors Authors | Bangaru S / Antanasijevic A / Ward A / Jackson AM | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structural mapping of antibody landscapes to human betacoronavirus spike proteins. Authors: Sandhya Bangaru / Aleksandar Antanasijevic / Nurgun Kose / Leigh M Sewall / Abigail M Jackson / Naveenchandra Suryadevara / Xiaoyan Zhan / Jonathan L Torres / Jeffrey Copps / Alba Torrents ...Authors: Sandhya Bangaru / Aleksandar Antanasijevic / Nurgun Kose / Leigh M Sewall / Abigail M Jackson / Naveenchandra Suryadevara / Xiaoyan Zhan / Jonathan L Torres / Jeffrey Copps / Alba Torrents de la Peña / James E Crowe / Andrew B Ward /  Abstract: Preexisting immunity against seasonal coronaviruses (CoVs) represents an important variable in predicting antibody responses and disease severity to severe acute respiratory syndrome CoV-2 (SARS-CoV- ...Preexisting immunity against seasonal coronaviruses (CoVs) represents an important variable in predicting antibody responses and disease severity to severe acute respiratory syndrome CoV-2 (SARS-CoV-2) infections. We used electron microscopy-based polyclonal epitope mapping (EMPEM) to characterize the antibody specificities against β-CoV spike proteins in prepandemic (PP) sera or SARS-CoV-2 convalescent (SC) sera. We observed that most PP sera had antibodies specific to seasonal human CoVs (HCoVs) OC43 and HKU1 spike proteins while the SC sera showed reactivity across all human β-CoVs. Detailed molecular mapping of spike-antibody complexes revealed epitopes that were differentially targeted by preexisting antibodies and SC serum antibodies. Our studies provide an antigenic landscape to β-HCoV spikes in the general population serving as a basis for cross-reactive epitope analyses in SARS-CoV-2-infected individuals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24969.map.gz emd_24969.map.gz | 139.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24969-v30.xml emd-24969-v30.xml emd-24969.xml emd-24969.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24969_fsc.xml emd_24969_fsc.xml | 12.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_24969.png emd_24969.png | 107.6 KB | ||
| Masks |  emd_24969_msk_1.map emd_24969_msk_1.map | 149.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-24969.cif.gz emd-24969.cif.gz | 7.3 KB | ||
| Others |  emd_24969_half_map_1.map.gz emd_24969_half_map_1.map.gz emd_24969_half_map_2.map.gz emd_24969_half_map_2.map.gz | 118.2 MB 118.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24969 http://ftp.pdbj.org/pub/emdb/structures/EMD-24969 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24969 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24969 | HTTPS FTP |
-Related structure data
| Related structure data |  7sb4MC  7sb3C  7sb5C  7sbvC  7sbwC  7sbxC  7sbyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24969.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24969.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_24969_msk_1.map emd_24969_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_24969_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_24969_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412)
| Entire | Name: Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412) |
|---|---|
| Components |
|
-Supramolecule #1: Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412)
| Supramolecule | Name: Structure of OC43 spike in complex with polyclonal Fab2 (Donor 1412) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Human coronavirus OC43 Human coronavirus OC43 |
-Macromolecule #1: Spike protein
| Macromolecule | Name: Spike protein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human coronavirus OC43 Human coronavirus OC43 |
| Molecular weight | Theoretical: 151.119891 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFLILLISLP TAFAVIGDLK CPLDSRTGSL NNIDTGPPSI STATVDVTNG LGTYYVLDRV YLNTTLFLNG YYPTSGSTYR NMALKGTDK LSTLWFKPPF LSDFINGIFA KVKNTKVFKD GVMYSEFPAI TIGSTFVNTS YSVVVQPRTI NSTQDGVNKL Q GLLEVSVC ...String: MFLILLISLP TAFAVIGDLK CPLDSRTGSL NNIDTGPPSI STATVDVTNG LGTYYVLDRV YLNTTLFLNG YYPTSGSTYR NMALKGTDK LSTLWFKPPF LSDFINGIFA KVKNTKVFKD GVMYSEFPAI TIGSTFVNTS YSVVVQPRTI NSTQDGVNKL Q GLLEVSVC QYNMCEYPHT ICHPKLGNHF KELWHMDTGV VSCLYKRNFT YDVNATYLYF HFYQEGGTFY AYFTDTGVVT KF LFNVYLG MALSHYYVMP LTCISRRDIG FTLEYWVTPL TSRQYLLAFN QDGIIFNAVD CMSDFMSEIK CKTQSIAPPT GVY ELNGYT VQPIADVYRR KPDLPNCNIE AWLNDKSVPS PLNWERKTFS NCNFNMSSLM SFIQADSFTC NNIDAAKIYG MCFS SITID KFAIPNGRKV DLQLGNLGYL QSFNYRIDTT ATSCQLYYNL PAANVSVSRF NPSTWNKRFG FIENSVFKPQ PAGVL TNHD VVYAQHCFKA PKNFCPCKLN SSLCVGSGPG KNNGIGTCPA GTNYLTCHNL CNPDPITFTG PYKCPQTKSL VGIGEH CSG LAVKSDYCGG NPCTCQPQAF LGWSADSCLQ GDKCNIFANL ILHDVNSGLT CSTDLQKANT DIKLGVCVNY DLYGISG QG IFVEVNATYY NSWQNLLYDS NGNLYGFRDY ITNRTFMIRS CYSGRVSAAF HANSSEPALL FRNIKCNYVF NNSLIRQL Q PINYFDSYLG CVVNAYNSTA ISVQTCDLTV GSGYCVDYSK NRRSRRAITT GYRFTNFEPF TVNSVNDSLE PVGGLYEIQ IPSEFTIGNM EEFIQTSSPK VTIDCAAFVC GDYAACKSQL VEYGSFCDNI NAILTEVNEL LDTTQLQVAN SLMNGVTLST KLKDGVNFN VDDINFSSVL GCLGSECSKA SSRSAIEDLL FDKVKLSDVG FVAAYNNCTG GAEIRDLICV QSYKGIKVLP P LLSENQIS GYTLAATSAS LFPPWTAAAG VPFYLNVQYR INGLGVTMDV LSQNQKLIAN AFNNALDAIQ EGFDATNSAL VK IQAVVNA NAEALNNLLQ QLSNRFGAIS SSLQEILSRL DPPEAEAQID RLINGRLTAL NAYVSQQLSD STLVKFSAAQ AME KVNECV KSQSSRINFC GNGNHIISLV QNAPYGLYFI HFSYVPTKYV TAKVSPGLCI AGDRGIAPKS GYFVNVNNTW MYTG SGYYY PEPITENNVV VMSTCAVNYT KAPYVMLNTS TPNLPDFREE LDQWFKNQTS VAPDLSLDYI NVTFLDLQVE MNRLQ EAIK VLNGSGYIPE APRDGQAYVR KDGEWVLLST FLGRSLEVLF QGPGHHHHHH HHSAWSHPQF EKGGGSGGGG SGGSAW SHP QFEK UniProtKB: Spike glycoprotein |
-Macromolecule #2: Human polyclonal Fab model with polyalanine backbone - Heavy chain
| Macromolecule | Name: Human polyclonal Fab model with polyalanine backbone - Heavy chain type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 9.634868 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 31 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #6: Sapienic acid
| Macromolecule | Name: Sapienic acid / type: ligand / ID: 6 / Number of copies: 3 / Formula: 8Z9 |
|---|---|
| Molecular weight | Theoretical: 254.408 Da |
| Chemical component information |  ChemComp-8Z9: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 / Component:
| ||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 7 sec. | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 2321 / Average exposure time: 10.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.3 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)