+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22707 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | BMV VLP assembled on oligoB with D6 symmetry | ||||||||||||
 Map data Map data | BMV VLP assembled on short oligo | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |   Brome mosaic virus Brome mosaic virus | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | ||||||||||||
 Authors Authors | Bond KM / Tsvetkova IB / Wang JC-Y / Jarrold MF / Dragnea B | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Small / Year: 2020 Journal: Small / Year: 2020Title: Virus Assembly Pathways: Straying Away but Not Too Far. Authors: Kevin Bond / Irina B Tsvetkova / Joseph Che-Yen Wang / Martin F Jarrold / Bogdan Dragnea /  Abstract: Non-enveloped RNA viruses pervade all domains of life. In a cell, they co-assemble from viral RNA and capsid proteins. Virus-like particles can form in vitro where virtually any non-cognate ...Non-enveloped RNA viruses pervade all domains of life. In a cell, they co-assemble from viral RNA and capsid proteins. Virus-like particles can form in vitro where virtually any non-cognate polyanionic cargo can be packaged. How only viral RNA gets selected for packaging in vivo, in presence of myriad other polyanionic species, has been a puzzle. Through a combination of charge detection mass spectrometry and cryo-electron microscopy, it is determined that co-assembling brome mosaic virus (BMV) coat proteins and nucleic acid oligomers results in capsid structures and stoichiometries that differ from the icosahedral virion. These previously unknown shell structures are strained and less stable than the native one. However, they contain large native structure fragments that can be recycled to form BMV virions, should a viral genome become available. The existence of such structures suggest the possibility of a previously unknown regulatory pathway for the packaging process inside cells. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22707.map.gz emd_22707.map.gz | 60 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22707-v30.xml emd-22707-v30.xml emd-22707.xml emd-22707.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22707.png emd_22707.png | 81.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22707 http://ftp.pdbj.org/pub/emdb/structures/EMD-22707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22707 | HTTPS FTP |
-Validation report
| Summary document |  emd_22707_validation.pdf.gz emd_22707_validation.pdf.gz | 386.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22707_full_validation.pdf.gz emd_22707_full_validation.pdf.gz | 386 KB | Display | |
| Data in XML |  emd_22707_validation.xml.gz emd_22707_validation.xml.gz | 6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22707 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22707 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22707 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22707 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_22707.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22707.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BMV VLP assembled on short oligo | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.68 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
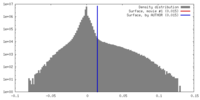
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Brome mosaic virus
| Entire | Name:   Brome mosaic virus Brome mosaic virus |
|---|---|
| Components |
|
-Supramolecule #1: Brome mosaic virus
| Supramolecule | Name: Brome mosaic virus / type: virus / ID: 1 / Parent: 0 Details: The purified BMV was disassembled and the RNA precipitated. The capsid protein was then assembled with 52 nt from a T-like region of the BMV RNA. NCBI-ID: 12302 / Sci species name: Brome mosaic virus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Experimental: 2.5 MDa |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 220.0 Å |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 4.5 / Component - Concentration: 100.0 mM / Component - Name: Ammonium acetate Details: 100 mM ammonium acetate that was pH adjusted to 4.5 with acetic acid |
|---|---|
| Grid | Model: C-flat / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: 15 s blotting time, 0 blotting force, and 100% humidity in Mark IV.. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)