+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21257 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J | |||||||||||||||||||||
 Map data Map data | BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | HIV / Rhesus macaque / antibody / Fab / vaccine / VIRAL PROTEIN | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | |||||||||||||||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.88 Å | |||||||||||||||||||||
 Authors Authors | Cottrell CA / Ward AB / Patel R | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2020 Journal: PLoS Pathog / Year: 2020Title: Mapping the immunogenic landscape of near-native HIV-1 envelope trimers in non-human primates. Authors: Christopher A Cottrell / Jelle van Schooten / Charles A Bowman / Meng Yuan / David Oyen / Mia Shin / Robert Morpurgo / Patricia van der Woude / Mariëlle van Breemen / Jonathan L Torres / ...Authors: Christopher A Cottrell / Jelle van Schooten / Charles A Bowman / Meng Yuan / David Oyen / Mia Shin / Robert Morpurgo / Patricia van der Woude / Mariëlle van Breemen / Jonathan L Torres / Raj Patel / Justin Gross / Leigh M Sewall / Jeffrey Copps / Gabriel Ozorowski / Bartek Nogal / Devin Sok / Eva G Rakasz / Celia Labranche / Vladimir Vigdorovich / Scott Christley / Diane G Carnathan / D Noah Sather / David Montefiori / Guido Silvestri / Dennis R Burton / John P Moore / Ian A Wilson / Rogier W Sanders / Andrew B Ward / Marit J van Gils /   Abstract: The induction of broad and potent immunity by vaccines is the key focus of research efforts aimed at protecting against HIV-1 infection. Soluble native-like HIV-1 envelope glycoproteins have shown ...The induction of broad and potent immunity by vaccines is the key focus of research efforts aimed at protecting against HIV-1 infection. Soluble native-like HIV-1 envelope glycoproteins have shown promise as vaccine candidates as they can induce potent autologous neutralizing responses in rabbits and non-human primates. In this study, monoclonal antibodies were isolated and characterized from rhesus macaques immunized with the BG505 SOSIP.664 trimer to better understand vaccine-induced antibody responses. Our studies reveal a diverse landscape of antibodies recognizing immunodominant strain-specific epitopes and non-neutralizing neo-epitopes. Additionally, we isolated a subset of mAbs against an epitope cluster at the gp120-gp41 interface that recognize the highly conserved fusion peptide and the glycan at position 88 and have characteristics akin to several human-derived broadly neutralizing antibodies. #1:  Journal: Biorxiv / Year: 2020 Journal: Biorxiv / Year: 2020Title: Mapping the immunogenic landscape of near-native HIV-1 envelope trimers in non-human primates Authors: Cottrell CA / van Schooten J / Bowman CA / Yuan M / Oyen D / Shin M / Morpurgo R / van der Woude P / van Breemen M / Torres JL / Patel R / Gross J / Sewall LM / Copps J / Ozorowski G / Nogal ...Authors: Cottrell CA / van Schooten J / Bowman CA / Yuan M / Oyen D / Shin M / Morpurgo R / van der Woude P / van Breemen M / Torres JL / Patel R / Gross J / Sewall LM / Copps J / Ozorowski G / Nogal B / Sok D / Rakasz EG / Labranche C / Vigdorovich V / Christley S / Carnathan DG / Sather DN / Montefiori D / Silvestri G / Burton DR / Moore JP / Wilson IA / Sanders RW / Ward AB / van Gils MJ | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21257.map.gz emd_21257.map.gz | 86 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21257-v30.xml emd-21257-v30.xml emd-21257.xml emd-21257.xml | 25 KB 25 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21257_fsc.xml emd_21257_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_21257.png emd_21257.png | 115.2 KB | ||
| Filedesc metadata |  emd-21257.cif.gz emd-21257.cif.gz | 7 KB | ||
| Others |  emd_21257_half_map_1.map.gz emd_21257_half_map_1.map.gz emd_21257_half_map_2.map.gz emd_21257_half_map_2.map.gz | 84.7 MB 84.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21257 http://ftp.pdbj.org/pub/emdb/structures/EMD-21257 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21257 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21257 | HTTPS FTP |
-Related structure data
| Related structure data |  6vo1MC  6vlrC  6vn0C  6vorC  6vosC  6vsrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21257.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21257.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
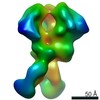
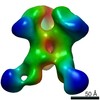
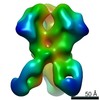
| Annotation | BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J
| File | emd_21257_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J
| File | emd_21257_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J
| Entire | Name: BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J |
|---|---|
| Components |
|
-Supramolecule #1: BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J
| Supramolecule | Name: BG505 SOSIP.v5.2 in complex with rhesus macaque Fab RM20J type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: BG505 SOSIP.v5.2
| Supramolecule | Name: BG505 SOSIP.v5.2 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #3: Fab RM20J
| Supramolecule | Name: Fab RM20J / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 53.268371 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETKKHNVWA THCCVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETKKHNVWA THCCVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT SAITQACPKV SFEPIPIHYC APAGFAILKC KDKKFNGTGP CPSVSTVQCT HGIKPVVSTQ LLLNGSLAEE EV MIRSENI TNNAKNILVQ FNTPVQINCT RPNNNTRKSI RIGPGQWFYA TGDIIGDIRQ AHCNVSKATW NETLGKVVKQ LRK HFGNNT IIRFANSSGG DLEVTTHSFN CGGEFFYCNT SGLFNSTWIS NTSVQGSNST GSNDSITLPC RIKQIINMWQ RIGQ AMYAP PIQGVIRCVS NITGLILTRD GGSTNSTTET FRPGGGDMRD NWRSELYKYK VVKIEPLGVA PTRCKRRVVG UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.178549 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEC QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #3: RM20J Heavy chain Fab
| Macromolecule | Name: RM20J Heavy chain Fab / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.714083 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPA VVQPSETLSL TCAVSGGSIS GGYGWTWIRQ APGKALEWIG NIYGHSGSTN YKSSLKRRLT ISTDTSKNQF SLKLTSVTA ADTAVYYCAR WSTADFDYWG QGVLVTVSS |
-Macromolecule #4: RM20J Kappa light chain
| Macromolecule | Name: RM20J Kappa light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.552784 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIVMTQSPSS LSASVGDTVT ITCRASQDIT NDLAWYQQKP GKAPKALIYY ASNLESGVPS RFSGSGAGTD FTLTISSLQP EDFALYYCQ QHNNYPLTFG PGTKVDIK |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 18 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6vo1: |
 Movie
Movie Controller
Controller









































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)