[English] 日本語
 Yorodumi
Yorodumi- EMDB-21021: MicroED structure of an oxindole product from modified trpB at 0.... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21021 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MicroED structure of an oxindole product from modified trpB at 0.9A resolution | |||||||||
 Map data Map data | MicroED 2Fo-Fc map for an oxindole product from modified trpB at 0.9A resolution | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | electron crystallography / cryo EM / Resolution: 0.9 Å | |||||||||
 Authors Authors | Dick M / Sarai NS / Martynowycz MW / Gonen T / Arnold FH | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Am Chem Soc / Year: 2019 Journal: J Am Chem Soc / Year: 2019Title: Tailoring Tryptophan Synthase TrpB for Selective Quaternary Carbon Bond Formation. Authors: Markus Dick / Nicholas S Sarai / Michael W Martynowycz / Tamir Gonen / Frances H Arnold /  Abstract: We previously engineered the β-subunit of tryptophan synthase (TrpB), which catalyzes the condensation of l-serine and indole to l-tryptophan, to synthesize a range of noncanonical amino acids from ...We previously engineered the β-subunit of tryptophan synthase (TrpB), which catalyzes the condensation of l-serine and indole to l-tryptophan, to synthesize a range of noncanonical amino acids from l-serine and indole derivatives or other nucleophiles. Here we employ directed evolution to engineer TrpB to accept 3-substituted oxindoles and form C-C bonds leading to new quaternary stereocenters. Initially, the variants that could use 3-substituted oxindoles preferentially formed N-C bonds on N of the substrate. Protecting N encouraged evolution toward C-alkylation, which persisted when protection was removed. Six generations of directed evolution resulted in TrpB with a 400-fold improvement in activity for alkylation of 3-substituted oxindoles and the ability to selectively form a new, all-carbon quaternary stereocenter at the γ-position of the amino acid products. The enzyme can also alkylate and form all-carbon quaternary stereocenters on structurally similar lactones and ketones, where it exhibits excellent regioselectivity for the tertiary carbon. The configurations of the γ-stereocenters of two of the products were determined via microcrystal electron diffraction (MicroED), and we report the MicroED structure of a small molecule obtained using the Falcon III direct electron detector. Highly thermostable and expressed at >500 mg/L culture, TrpB offers an efficient, sustainable, and selective platform for the construction of diverse noncanonical amino acids bearing all-carbon quaternary stereocenters. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21021.map.gz emd_21021.map.gz | 558 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21021-v30.xml emd-21021-v30.xml emd-21021.xml emd-21021.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21021.png emd_21021.png | 43.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21021 http://ftp.pdbj.org/pub/emdb/structures/EMD-21021 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21021 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21021 | HTTPS FTP |
-Validation report
| Summary document |  emd_21021_validation.pdf.gz emd_21021_validation.pdf.gz | 79.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21021_full_validation.pdf.gz emd_21021_full_validation.pdf.gz | 78.8 KB | Display | |
| Data in XML |  emd_21021_validation.xml.gz emd_21021_validation.xml.gz | 492 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21021 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21021 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21021 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21021 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21021.map.gz / Format: CCP4 / Size: 4.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21021.map.gz / Format: CCP4 / Size: 4.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | MicroED 2Fo-Fc map for an oxindole product from modified trpB at 0.9A resolution | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X: 0.28929 Å / Y: 0.28063 Å / Z: 0.28556 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
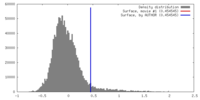
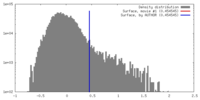
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : (S)-2-amino-3-((S)-3-methyl-2-oxoindolin-3-yl)propanoic acid
| Entire | Name: (S)-2-amino-3-((S)-3-methyl-2-oxoindolin-3-yl)propanoic acid |
|---|---|
| Components |
|
-Supramolecule #1: (S)-2-amino-3-((S)-3-methyl-2-oxoindolin-3-yl)propanoic acid
| Supramolecule | Name: (S)-2-amino-3-((S)-3-methyl-2-oxoindolin-3-yl)propanoic acid type: complex / ID: 1 / Parent: 0 / Details: Oxindole product from TrpB |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 215 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron crystallography |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI CETA (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Sampling interval: 28.0 µm / Number grids imaged: 2 / Number real images: 120 / Number diffraction images: 120 / Average exposure time: 1.0 sec. / Average electron dose: 0.01 e/Å2 Details: Frames were recorded every 0.5 degrees for ~60 degrees with a 1 second integration time. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: DIFFRACTION / Camera length: 1053 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN / Tilt angle: 0, 60 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 0.9 Å / Resolution method: DIFFRACTION PATTERN/LAYERLINES |
|---|---|
| Crystal parameters | Unit cell - A: 12.15 Å / Unit cell - B: 8.98 Å / Unit cell - C: 10.28 Å / Unit cell - γ: 90 ° / Unit cell - α: 90 ° / Unit cell - β: 99.41 ° / Space group: 14 |
| Crystallography statistics | Number intensities measured: 2524 / Number structure factors: 1028 / Fourier space coverage: 65.1 / R sym: 12.8 / R merge: 10.1 / Overall phase error: 20 / Overall phase residual: 20 / Phase error rejection criteria: 0 / High resolution: 0.9 Å / Shell - Shell ID: 1 / Shell - High resolution: 10.0 Å / Shell - Low resolution: 0.9 Å / Shell - Number structure factors: 1024 / Shell - Phase residual: 20 / Shell - Fourier space coverage: 65.1 / Shell - Multiplicity: 2.4 |
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL / Overall B value: 6 / Target criteria: Least Squares |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)