+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20697 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
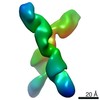
| Title | RNA X with 24 PTX | |||||||||
 Map data Map data | RNA X with 24 PTX | |||||||||
 Sample Sample |
| |||||||||
| Biological species | unidentified (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.0 Å | |||||||||
 Authors Authors | Zhang K / Guo S / Chiu W | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Ultra-thermostable RNA nanoparticles for solubilizing and high-yield loading of paclitaxel for breast cancer therapy. Authors: Sijin Guo / Mario Vieweger / Kaiming Zhang / Hongran Yin / Hongzhi Wang / Xin Li / Shanshan Li / Shuiying Hu / Alex Sparreboom / B Mark Evers / Yizhou Dong / Wah Chiu / Peixuan Guo /  Abstract: Paclitaxel is widely used in cancer treatments, but poor water-solubility and toxicity raise serious concerns. Here we report an RNA four-way junction nanoparticle with ultra-thermodynamic stability ...Paclitaxel is widely used in cancer treatments, but poor water-solubility and toxicity raise serious concerns. Here we report an RNA four-way junction nanoparticle with ultra-thermodynamic stability to solubilize and load paclitaxel for targeted cancer therapy. Each RNA nanoparticle covalently loads twenty-four paclitaxel molecules as a prodrug. The RNA-paclitaxel complex is structurally rigid and stable, demonstrated by the sub-nanometer resolution imaging of cryo-EM. Using RNA nanoparticles as carriers increases the water-solubility of paclitaxel by 32,000-fold. Intravenous injections of RNA-paclitaxel nanoparticles with specific cancer-targeting ligand dramatically inhibit breast cancer growth, with nearly undetectable toxicity and immune responses in mice. No fatalities are observed at a paclitaxel dose equal to the reported LD. The use of ultra-thermostable RNA nanoparticles to deliver chemical prodrugs addresses issues with RNA unfolding and nanoparticle dissociation after high-density drug loading. This finding provides a stable nano-platform for chemo-drug delivery as well as an efficient method to solubilize hydrophobic drugs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20697.map.gz emd_20697.map.gz | 39.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20697-v30.xml emd-20697-v30.xml emd-20697.xml emd-20697.xml | 10.6 KB 10.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20697.png emd_20697.png | 93.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20697 http://ftp.pdbj.org/pub/emdb/structures/EMD-20697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20697 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20697 | HTTPS FTP |
-Validation report
| Summary document |  emd_20697_validation.pdf.gz emd_20697_validation.pdf.gz | 78.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20697_full_validation.pdf.gz emd_20697_full_validation.pdf.gz | 77.8 KB | Display | |
| Data in XML |  emd_20697_validation.xml.gz emd_20697_validation.xml.gz | 495 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20697 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20697 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20697 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20697 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20697.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20697.map.gz / Format: CCP4 / Size: 42.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RNA X with 24 PTX | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Designed RNA X nanoparticle with 24 PTX (paclitaxel)
| Entire | Name: Designed RNA X nanoparticle with 24 PTX (paclitaxel) |
|---|---|
| Components |
|
-Supramolecule #1: Designed RNA X nanoparticle with 24 PTX (paclitaxel)
| Supramolecule | Name: Designed RNA X nanoparticle with 24 PTX (paclitaxel) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Experimental: 60 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-30 / Number grids imaged: 1 / Number real images: 3800 / Average exposure time: 6.0 sec. / Average electron dose: 45.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















