[English] 日本語
 Yorodumi
Yorodumi- EMDB-20336: Cryo-electron tomogram of Bacillus subtilis wild type sporangium ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20336 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
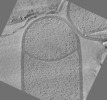
| Title | Cryo-electron tomogram of Bacillus subtilis wild type sporangium (curved septum) shown in Figure 1F of the publication | |||||||||
 Map data Map data | Cryo-electron tomogram of Bacillus subtilis wild type sporangium with curved septum | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Khanna K / Villa E | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: The molecular architecture of engulfment during sporulation. Authors: Kanika Khanna / Javier Lopez-Garrido / Ziyi Zhao / Reika Watanabe / Yuan Yuan / Joseph Sugie / Kit Pogliano / Elizabeth Villa /  Abstract: The study of bacterial cell biology is limited by difficulties in visualizing cellular structures at high spatial resolution within their native milieu. Here, we visualize sporulation using cryo- ...The study of bacterial cell biology is limited by difficulties in visualizing cellular structures at high spatial resolution within their native milieu. Here, we visualize sporulation using cryo-electron tomography coupled with cryo-focused ion beam milling, allowing the reconstruction of native-state cellular sections at molecular resolution. During sporulation, an asymmetrically-positioned septum generates a larger mother cell and a smaller forespore. Subsequently, the mother cell engulfs the forespore. We show that the septal peptidoglycan is not completely degraded at the onset of engulfment. Instead, the septum is uniformly and only slightly thinned as it curves towards the mother cell. Then, the mother cell membrane migrates around the forespore in tiny finger-like projections, whose formation requires the mother cell SpoIIDMP protein complex. We propose that a limited number of SpoIIDMP complexes tether to and degrade the peptidoglycan ahead of the engulfing membrane, generating an irregular membrane front. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20336.map.gz emd_20336.map.gz | 24.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20336-v30.xml emd-20336-v30.xml emd-20336.xml emd-20336.xml | 8.8 KB 8.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20336.png emd_20336.png | 132.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20336 http://ftp.pdbj.org/pub/emdb/structures/EMD-20336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20336 | HTTPS FTP |
-Validation report
| Summary document |  emd_20336_validation.pdf.gz emd_20336_validation.pdf.gz | 78.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20336_full_validation.pdf.gz emd_20336_full_validation.pdf.gz | 77.3 KB | Display | |
| Data in XML |  emd_20336_validation.xml.gz emd_20336_validation.xml.gz | 500 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20336 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20336 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20336 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20336 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20336.map.gz / Format: CCP4 / Size: 37.5 MB / Type: IMAGE STORED AS SIGNED BYTE Download / File: emd_20336.map.gz / Format: CCP4 / Size: 37.5 MB / Type: IMAGE STORED AS SIGNED BYTE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-electron tomogram of Bacillus subtilis wild type sporangium with curved septum | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 24.48 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacillus subtilis wild type sporangium with flat septum
| Entire | Name: Bacillus subtilis wild type sporangium with flat septum |
|---|---|
| Components |
|
-Supramolecule #1: Bacillus subtilis wild type sporangium with flat septum
| Supramolecule | Name: Bacillus subtilis wild type sporangium with flat septum type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE-PROPANE |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 kV / Focused ion beam - Current: 0.03 nA / Focused ion beam - Duration: 3600 sec. / Focused ion beam - Temperature: 100 K / Focused ion beam - Initial thickness: 1200 nm / Focused ion beam - Final thickness: 200 nm Focused ion beam - Details: The value given for _emd_sectioning_focused_ion_beam.instrument is FEI Scios Dual Beam. This is not in a list of allowed values set(['DB235', 'OTHER']) so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Software - Name:  IMOD / Number images used: 68 IMOD / Number images used: 68 |
|---|
 Movie
Movie Controller
Controller