[English] 日本語
 Yorodumi
Yorodumi- EMDB-20001: Structure of canine parvovirus in complex with transferrin recept... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20001 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
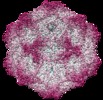
| Title | Structure of canine parvovirus in complex with transferrin receptor type-1 | |||||||||
 Map data Map data | Icosahedral 3D map of CPV incubated with TfR | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CPV / TfR / icosahedral / cryo / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont entry into host cell via permeabilization of host membrane / microtubule-dependent intracellular transport of viral material towards nucleus / adhesion receptor-mediated virion attachment to host cell / T=1 icosahedral viral capsid / viral penetration into host nucleus / host cell / clathrin-dependent endocytosis of virus by host cell / entry receptor-mediated virion attachment to host cell / host cell nucleus / structural molecule activity / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Canine parvovirus 2 Canine parvovirus 2 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Lee H / Hafenstein S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: Transferrin receptor binds virus capsid with dynamic motion. Authors: Hyunwook Lee / Heather M Callaway / Javier O Cifuente / Carol M Bator / Colin R Parrish / Susan L Hafenstein /   Abstract: Canine parvovirus (CPV) is an important pathogen causing severe diseases in dogs, including acute hemorrhagic enteritis, myocarditis, and cerebellar disease. Cross-species transmission of CPV occurs ...Canine parvovirus (CPV) is an important pathogen causing severe diseases in dogs, including acute hemorrhagic enteritis, myocarditis, and cerebellar disease. Cross-species transmission of CPV occurs as a result of mutations on the viral capsid surface that alter the species-specific binding to the host receptor, transferrin receptor type-1 (TfR). The interaction between CPV and TfR has been extensively studied, and previous analyses have suggested that the CPV-TfR complex is asymmetric. To enhance the understanding of the underlying molecular mechanisms, we determined the CPV-TfR interaction using cryo-electron microscopy to solve the icosahedral (3.0-Å resolution) and asymmetric (5.0-Å resolution) complex structures. Structural analyses revealed conformational variations of the TfR molecules relative to the binding site, which translated into dynamic molecular interactions between CPV and TfR. The precise footprint of the receptor on the virus capsid was identified, along with the identity of the amino acid residues in the virus-receptor interface. Our "rock-and-roll" model provides an explanation for previous findings and gives insights into species jumping and the variation in host ranges associated with new pandemics in dogs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20001.map.gz emd_20001.map.gz | 227.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20001-v30.xml emd-20001-v30.xml emd-20001.xml emd-20001.xml | 9.8 KB 9.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20001.png emd_20001.png | 330.3 KB | ||
| Filedesc metadata |  emd-20001.cif.gz emd-20001.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20001 http://ftp.pdbj.org/pub/emdb/structures/EMD-20001 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20001 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20001 | HTTPS FTP |
-Related structure data
| Related structure data |  6oasMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20001.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20001.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral 3D map of CPV incubated with TfR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Canine parvovirus 2
| Entire | Name:  Canine parvovirus 2 Canine parvovirus 2 |
|---|---|
| Components |
|
-Supramolecule #1: Canine parvovirus 2
| Supramolecule | Name: Canine parvovirus 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 246878 / Sci species name: Canine parvovirus 2 / Virus type: VIRION / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Canine parvovirus 2 Canine parvovirus 2 |
| Molecular weight | Theoretical: 61.562367 KDa |
| Sequence | String: GVGISTGTFN NQTEFKFLEN GWVEITANSS RLVHLNMPES ENYRRVVVNN MDKTAVNGNM ALDDIHAQIV TPWSLVDANA WGVWFNPGD WQLIVNTMSE LHLVSFEQEI FNVVLKTVSE SATQPPTKVY NNDLTASLMV ALDSNNTMPF TPAAMRSETL G FYPWKPTI ...String: GVGISTGTFN NQTEFKFLEN GWVEITANSS RLVHLNMPES ENYRRVVVNN MDKTAVNGNM ALDDIHAQIV TPWSLVDANA WGVWFNPGD WQLIVNTMSE LHLVSFEQEI FNVVLKTVSE SATQPPTKVY NNDLTASLMV ALDSNNTMPF TPAAMRSETL G FYPWKPTI PTPWRYYFQW DRTLIPSHTG TSGTPTNIYH GTDPDDVQFY TIENSVPVHL LRTGDEFATG TFFFDCKPCR LT HTWQTNR ALGLPPFLNS LPQSEGATNF GDIGVQQDKR RGVTQMGNTN YITEATIMRP AEVGYSAPYY SFEASTQGPF KTP IAAGRG GAQTDENQAA DGNPRYAFGR QHGQKTTTTG ETPERFTYIA HQDTGRYPEG DWIQNINFNL PVTNDNVLLP TDPI GGKTG INYTNIFNTY GPLTALNNVP PVYPNGQIWD KEFDTDLKPR LHVNAPFVCQ NNCPGQLFVK VAPNLTNEYD PDASA NMSR IVTYSDFWWK GKLVFKAKLR ASHTWNPIQQ MSINVDNQFN YVPSNIGGMK IVYEKSQLAP RKLY UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 150.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 3.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 132120 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















