+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human RNF213: focused refinement of E3 ligase domain | |||||||||
 Map data Map data | Unsharpened map of RNF213 E3 ligase domain after focused refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | E3 ubiquitin ligase / ANTIMICROBIAL PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
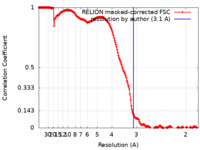
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Naydenova K / Randow F | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: EMBO Rep / Year: 2024 Journal: EMBO Rep / Year: 2024Title: Recognition of phylogenetically diverse pathogens through enzymatically amplified recruitment of RNF213. Authors: Ana Crespillo-Casado / Prathyush Pothukuchi / Katerina Naydenova / Matthew C J Yip / Janet M Young / Jerome Boulanger / Vimisha Dharamdasani / Ceara Harper / Pierre-Mehdi Hammoudi / Elsje G ...Authors: Ana Crespillo-Casado / Prathyush Pothukuchi / Katerina Naydenova / Matthew C J Yip / Janet M Young / Jerome Boulanger / Vimisha Dharamdasani / Ceara Harper / Pierre-Mehdi Hammoudi / Elsje G Otten / Keith Boyle / Mayuri Gogoi / Harmit S Malik / Felix Randow /   Abstract: Innate immunity senses microbial ligands known as pathogen-associated molecular patterns (PAMPs). Except for nucleic acids, PAMPs are exceedingly taxa-specific, thus enabling pattern recognition ...Innate immunity senses microbial ligands known as pathogen-associated molecular patterns (PAMPs). Except for nucleic acids, PAMPs are exceedingly taxa-specific, thus enabling pattern recognition receptors to detect cognate pathogens while ignoring others. How the E3 ubiquitin ligase RNF213 can respond to phylogenetically distant pathogens, including Gram-negative Salmonella, Gram-positive Listeria, and eukaryotic Toxoplasma, remains unknown. Here we report that the evolutionary history of RNF213 is indicative of repeated adaptation to diverse pathogen target structures, especially in and around its newly identified CBM20 carbohydrate-binding domain, which we have resolved by cryo-EM. We find that RNF213 forms coats on phylogenetically distant pathogens. ATP hydrolysis by RNF213's dynein-like domain is essential for coat formation on all three pathogens studied as is RZ finger-mediated E3 ligase activity for bacteria. Coat formation is not diffusion-limited but instead relies on rate-limiting initiation events and subsequent cooperative incorporation of further RNF213 molecules. We conclude that RNF213 responds to evolutionarily distant pathogens through enzymatically amplified cooperative recruitment. #1:  Journal: Biorxiv / Year: 2024 Journal: Biorxiv / Year: 2024Title: Recognition of phylogenetically diverse pathogens through enzymatically amplified recruitment of RNF213 Authors: Casado AC / Pothukuchi P / Naydenova K / Yip MCJ / Young JM / Boulanger J / Dharamdasani V / Harper C / Hammoudi PM / Otten EG / Boyle K / Gogoi M / Malik HS / Randow F | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19658.map.gz emd_19658.map.gz | 475 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19658-v30.xml emd-19658-v30.xml emd-19658.xml emd-19658.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19658_fsc.xml emd_19658_fsc.xml | 18.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_19658.png emd_19658.png | 59.3 KB | ||
| Masks |  emd_19658_msk_1.map emd_19658_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19658.cif.gz emd-19658.cif.gz | 4.9 KB | ||
| Others |  emd_19658_half_map_1.map.gz emd_19658_half_map_1.map.gz emd_19658_half_map_2.map.gz emd_19658_half_map_2.map.gz | 413.4 MB 413.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19658 http://ftp.pdbj.org/pub/emdb/structures/EMD-19658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19658 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19658.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19658.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map of RNF213 E3 ligase domain after focused refinement | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.921 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19658_msk_1.map emd_19658_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
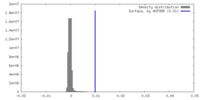
| Density Histograms |
-Half map: Half-map (2) of RNF213 E3 ligase domain focused refinement
| File | emd_19658_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map (2) of RNF213 E3 ligase domain focused refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
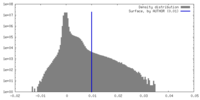
| Density Histograms |
-Half map: Half-map (1) of RNF213 E3 ligase domain focused refinement
| File | emd_19658_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map (1) of RNF213 E3 ligase domain focused refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
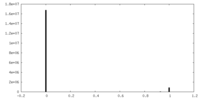
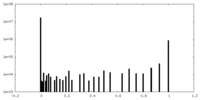
| Density Histograms |
- Sample components
Sample components
-Entire : Human RNF213
| Entire | Name: Human RNF213 |
|---|---|
| Components |
|
-Supramolecule #1: Human RNF213
| Supramolecule | Name: Human RNF213 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: E3 ubiquitin ligase RNF213, human, N1045D natural variant |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 590 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Average exposure time: 4.86 sec. / Average electron dose: 29.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Protocol: AB INITIO MODEL |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)