[English] 日本語
 Yorodumi
Yorodumi- EMDB-19481: Conformation-B of the full-length outer membrane core complex (Tr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Conformation-B of the full-length outer membrane core complex (TrwH/VirB7, TrwF/VirB9, TrwE/VirB10CTD) from the fully-assembled R388 type IV secretion system determined by cryo-EM. | |||||||||||||||
 Map data Map data | OMCC_Conformation-B _ Unsharpened | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | type IV secretion system type 4 secretion system T4SS OMCC Conformation-B core complex outer membrane complex R388 plasmid conjugation bacterial secretion secretion secretion system protein complex VirB10 VirB9 VirB7 TrwE TrwF TrwH / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||||||||
| Biological species |  | |||||||||||||||
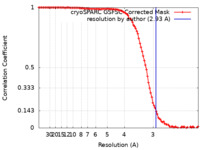
| Method | single particle reconstruction / cryo EM / Resolution: 2.93 Å | |||||||||||||||
 Authors Authors | Mace K / Waksman G | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: EMBO J / Year: 2024 Journal: EMBO J / Year: 2024Title: Cryo-EM structure of a conjugative type IV secretion system suggests a molecular switch regulating pilus biogenesis. Authors: Kévin Macé / Gabriel Waksman /   Abstract: Conjugative type IV secretion systems (T4SS) mediate bacterial conjugation, a process that enables the unidirectional exchange of genetic materials between a donor and a recipient bacterial cell. ...Conjugative type IV secretion systems (T4SS) mediate bacterial conjugation, a process that enables the unidirectional exchange of genetic materials between a donor and a recipient bacterial cell. Bacterial conjugation is the primary means by which antibiotic resistance genes spread among bacterial populations (Barlow 2009; Virolle et al, 2020). Conjugative T4SSs form pili: long extracellular filaments that connect with recipient cells. Previously, we solved the cryo-electron microscopy (cryo-EM) structure of a conjugative T4SS. In this article, based on additional data, we present a more complete T4SS cryo-EM structure than that published earlier. Novel structural features include details of the mismatch symmetry within the OMCC, the presence of a fourth VirB8 subunit in the asymmetric unit of both the arches and the inner membrane complex (IMC), and a hydrophobic VirB5 tip in the distal end of the stalk. Additionally, we provide previously undescribed structural insights into the protein VirB10 and identify a novel regulation mechanism of T4SS-mediated pilus biogenesis by this protein, that we believe is a key checkpoint for this process. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19481.map.gz emd_19481.map.gz | 50.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19481-v30.xml emd-19481-v30.xml emd-19481.xml emd-19481.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19481_fsc.xml emd_19481_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_19481.png emd_19481.png | 113.6 KB | ||
| Masks |  emd_19481_msk_1.map emd_19481_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19481.cif.gz emd-19481.cif.gz | 5.9 KB | ||
| Others |  emd_19481_additional_1.map.gz emd_19481_additional_1.map.gz emd_19481_half_map_1.map.gz emd_19481_half_map_1.map.gz emd_19481_half_map_2.map.gz emd_19481_half_map_2.map.gz | 91 MB 95.1 MB 95.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19481 http://ftp.pdbj.org/pub/emdb/structures/EMD-19481 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19481 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19481 | HTTPS FTP |
-Validation report
| Summary document |  emd_19481_validation.pdf.gz emd_19481_validation.pdf.gz | 915.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19481_full_validation.pdf.gz emd_19481_full_validation.pdf.gz | 915.2 KB | Display | |
| Data in XML |  emd_19481_validation.xml.gz emd_19481_validation.xml.gz | 18.2 KB | Display | |
| Data in CIF |  emd_19481_validation.cif.gz emd_19481_validation.cif.gz | 23.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19481 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19481 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19481 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19481 | HTTPS FTP |
-Related structure data
| Related structure data |  8rt7MC  8rt4C  8rt5C  8rt6C  8rt8C  8rt9C  8rtaC  8rtbC  8rtdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19481.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19481.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | OMCC_Conformation-B _ Unsharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.067 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19481_msk_1.map emd_19481_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: OMCC Conformation-B Sharpened
| File | emd_19481_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | OMCC_Conformation-B _ Sharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-A
| File | emd_19481_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-B
| File | emd_19481_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Conformation-B of the outer membrane core complex from the fully-...
| Entire | Name: Conformation-B of the outer membrane core complex from the fully-assembled R388 type IV secretion system |
|---|---|
| Components |
|
-Supramolecule #1: Conformation-B of the outer membrane core complex from the fully-...
| Supramolecule | Name: Conformation-B of the outer membrane core complex from the fully-assembled R388 type IV secretion system type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: TrwE protein
| Macromolecule | Name: TrwE protein / type: protein_or_peptide / ID: 1 / Details: Sequence from conjugative plasmid R388 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.443785 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MFGRKKGDVI DAGAELERAE QERIEGEYGA SELASERRPH TPGARTLLMV LLCVIAVVLV TLSYKAYKVR GVVEDDDAQP QQVVRQVIP GYTPRPIRPE PENVPEPPQP TTSVPAIQPA PVTQPVRPQP TGPREKTPYE LARERMLRSG LTAGSGGGED L PRPQGGDV ...String: MFGRKKGDVI DAGAELERAE QERIEGEYGA SELASERRPH TPGARTLLMV LLCVIAVVLV TLSYKAYKVR GVVEDDDAQP QQVVRQVIP GYTPRPIRPE PENVPEPPQP TTSVPAIQPA PVTQPVRPQP TGPREKTPYE LARERMLRSG LTAGSGGGED L PRPQGGDV PAGGLMGGGG GGGELAEKLQ PMRLSGSSAG RLGNRDMLIT QGTQLDCVLE TRLVTTQPGM TTCHLTRDVY ST SGRVVLL DRGSKVVGFY QGGLRQGQAR IFVQWSRIET PSGVVINLDS PGTGPLGEAG LGGWIDRHFW ERFGGAIMIS LIG DLGDWA SRQGSRQGDN SIQFSNTANG VESAAAEALR NSINIPPTLY KNQGERVNIL VARDLDFSDV YSLESIPTK UniProtKB: TrwE protein |
-Macromolecule #2: TrwF protein
| Macromolecule | Name: TrwF protein / type: protein_or_peptide / ID: 2 / Details: Sequence from conjugative plasmid R388 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.749586 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKLAIVALL ASLHAVPALA LDVPSSSRYD HRIRYVTYNP ADVVQVDTVL GVATHIMLEE GEQYLTHAFG DSEAYAFARK GRHIFIKPQ AELANTNLIV VTDRRSYKFR LQMRNDRNGA MYELAFRYPD TQARQTREAN ARAAVEAAFE QRVGAYYNLK Y MMSGDKDI ...String: MKKLAIVALL ASLHAVPALA LDVPSSSRYD HRIRYVTYNP ADVVQVDTVL GVATHIMLEE GEQYLTHAFG DSEAYAFARK GRHIFIKPQ AELANTNLIV VTDRRSYKFR LQMRNDRNGA MYELAFRYPD TQARQTREAN ARAAVEAAFE QRVGAYYNLK Y MMSGDKDI APVNAWDDGR FTYFKFSANA DLPSIYFVDA EGNESLVPRT TVGSSNNIIA VHKVNPKWMI RLGNRALAIF NE AYDPNGV PNDTGTASPA VRRVNKGGN UniProtKB: TrwF protein |
-Macromolecule #3: TrwH
| Macromolecule | Name: TrwH / type: protein_or_peptide / ID: 3 / Details: Sequence from conjugative plasmid R388 / Number of copies: 14 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5.089048 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIFAILM TGLLSACASA PKPKQPSDFN REPVNKTVPV EIQRGAL UniProtKB: TrwH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 3.3000000000000003 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)