[English] 日本語
 Yorodumi
Yorodumi- EMDB-18793: Cryo-EM density map of DNMT3A1-DNMT3L on a human H2AKc119ub nucle... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM density map of DNMT3A1-DNMT3L on a human H2AKc119ub nucleosome at 5.1 A resolution | |||||||||
 Map data Map data | DNMT3A1-DNMT3L_H2AKc119ub_unmaskedmap1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chromatin / Nucleosome / methyltransferase / Ubiquitin / DNA BINDING PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
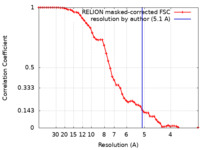
| Method | single particle reconstruction / cryo EM / Resolution: 5.1 Å | |||||||||
 Authors Authors | Wapenaar H / Wilson MD | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO Rep / Year: 2024 Journal: EMBO Rep / Year: 2024Title: The N-terminal region of DNMT3A engages the nucleosome surface to aid chromatin recruitment. Authors: Hannah Wapenaar / Gillian Clifford / Willow Rolls / Moira Pasquier / Hayden Burdett / Yujie Zhang / Gauri Deák / Juan Zou / Christos Spanos / Mark R D Taylor / Jacquie Mills / James A ...Authors: Hannah Wapenaar / Gillian Clifford / Willow Rolls / Moira Pasquier / Hayden Burdett / Yujie Zhang / Gauri Deák / Juan Zou / Christos Spanos / Mark R D Taylor / Jacquie Mills / James A Watson / Dhananjay Kumar / Richard Clark / Alakta Das / Devisree Valsakumar / Janice Bramham / Philipp Voigt / Duncan Sproul / Marcus D Wilson /  Abstract: DNA methyltransferase 3A (DNMT3A) plays a critical role in establishing and maintaining DNA methylation patterns in vertebrates. Here we structurally and biochemically explore the interaction of ...DNA methyltransferase 3A (DNMT3A) plays a critical role in establishing and maintaining DNA methylation patterns in vertebrates. Here we structurally and biochemically explore the interaction of DNMT3A1 with diverse modified nucleosomes indicative of different chromatin environments. A cryo-EM structure of the full-length DNMT3A1-DNMT3L complex with a H2AK119ub nucleosome reveals that the DNMT3A1 ubiquitin-dependent recruitment (UDR) motif interacts specifically with H2AK119ub and makes extensive contacts with the core nucleosome histone surface. This interaction facilitates robust DNMT3A1 binding to nucleosomes, and previously unexplained DNMT3A disease-associated mutations disrupt this interface. Furthermore, the UDR-nucleosome interaction synergises with other DNMT3A chromatin reading elements in the absence of histone ubiquitylation. H2AK119ub does not stimulate DNMT3A DNA methylation activity, as observed for the previously described H3K36me2 mark, which may explain low levels of DNA methylation on H2AK119ub marked facultative heterochromatin. This study highlights the importance of multivalent binding of DNMT3A to histone modifications and the nucleosome surface and increases our understanding of how DNMT3A1 chromatin recruitment occurs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18793.map.gz emd_18793.map.gz | 14.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18793-v30.xml emd-18793-v30.xml emd-18793.xml emd-18793.xml | 29.5 KB 29.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18793_fsc.xml emd_18793_fsc.xml | 5.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_18793.png emd_18793.png | 59.2 KB | ||
| Masks |  emd_18793_msk_1.map emd_18793_msk_1.map | 15.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18793.cif.gz emd-18793.cif.gz | 7.4 KB | ||
| Others |  emd_18793_additional_1.map.gz emd_18793_additional_1.map.gz emd_18793_half_map_1.map.gz emd_18793_half_map_1.map.gz emd_18793_half_map_2.map.gz emd_18793_half_map_2.map.gz | 3.5 MB 11.9 MB 11.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18793 http://ftp.pdbj.org/pub/emdb/structures/EMD-18793 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18793 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18793 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18793.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18793.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DNMT3A1-DNMT3L_H2AKc119ub_unmaskedmap1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.658 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18793_msk_1.map emd_18793_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Maskedmap1
| File | emd_18793_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Maskedmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap1 map1
| File | emd_18793_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap1_map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap2 map1
| File | emd_18793_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap2_map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : DNMT3A1-DNMT3L in complex with a human H2AKc119ub nucleosome wrap...
+Supramolecule #1: DNMT3A1-DNMT3L in complex with a human H2AKc119ub nucleosome wrap...
+Supramolecule #2: H2AKc119ub Nucleosome wrapped with 195 bp Widom601 DNA
+Supramolecule #3: DNMT3A1-DNMT3L complex
+Macromolecule #1: Histone H3
+Macromolecule #2: Histone H2AK119C
+Macromolecule #3: ubiquitin
+Macromolecule #4: Histone H4
+Macromolecule #6: DNMT3L
+Macromolecule #7: Histone H2B
+Macromolecule #8: DNMT3A1
+Macromolecule #9: 195bp Widom601 DNA
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.13 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 15 mM HEPES pH 7.5, 65 mM NaCl, 1 mM DTT, 0.1 mM S-Adenosyl methionine | |||||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.038 kPa | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 6969 / Average electron dose: 61.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)