+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1873 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
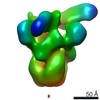
| Title | The flagellar cap, HAP2 pentamer | |||||||||
 Map data Map data | This is a HAP2 pentamer. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | flagellar cap /  self assembly / self assembly /  motor protein / motor protein /  flagellum flagellum | |||||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 26.0 Å cryo EM / Resolution: 26.0 Å | |||||||||
 Authors Authors | Maki-Yonekura S / Yonekura K / Namba K | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2003 Journal: Proc Natl Acad Sci U S A / Year: 2003Title: Domain movements of HAP2 in the cap-filament complex formation and growth process of the bacterial flagellum. Authors: Saori Maki-Yonekura / Koji Yonekura / Keiichi Namba /  Abstract: The cap at the growing end of the bacterial flagellum is essential for its growth, remaining stably attached while permitting the insertion of flagellin transported from the cytoplasm through the ...The cap at the growing end of the bacterial flagellum is essential for its growth, remaining stably attached while permitting the insertion of flagellin transported from the cytoplasm through the narrow central channel. We analyzed the structure of the isolated cap in its frozen hydrated state by electron cryomicroscopy. The 3D density map now shows detailed features of domains and their connections, giving reliable volumes and masses, making assignment of the domains to the amino acid sequence possible. A model of the cap-filament complex built with an atomic model of the filament allows a quantitative analysis of the cap domain movements on cap binding and rotation that promotes the efficient self assembly of flagellin during the filament growth process. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1873.map.gz emd_1873.map.gz | 448.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1873-v30.xml emd-1873-v30.xml emd-1873.xml emd-1873.xml | 6.5 KB 6.5 KB | Display Display |  EMDB header EMDB header |
| Images |  1873.png 1873.png | 64.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1873 http://ftp.pdbj.org/pub/emdb/structures/EMD-1873 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1873 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1873 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1873.map.gz / Format: CCP4 / Size: 475.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1873.map.gz / Format: CCP4 / Size: 475.6 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a HAP2 pentamer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.45 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Flagellar hook-associated protein 2, HAP2
| Entire | Name: Flagellar hook-associated protein 2, HAP2 |
|---|---|
| Components |
|
-Supramolecule #1000: Flagellar hook-associated protein 2, HAP2
| Supramolecule | Name: Flagellar hook-associated protein 2, HAP2 / type: sample / ID: 1000 / Details: The sample was monodisperse / Oligomeric state: Decamer / Number unique components: 1 |
|---|
-Macromolecule #1: Flagellar cap
| Macromolecule | Name: Flagellar cap / type: protein_or_peptide / ID: 1 / Name.synonym: Flagellar hook-associated protein 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) / Recombinant plasmid: pKOT134 Escherichia coli BL21(DE3) (bacteria) / Recombinant plasmid: pKOT134 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: Manual freezing |
|---|
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3000SFF |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Sample stage | Specimen holder: Top-entry / Specimen holder model: JEOL |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C5 (5 fold cyclic ) / Resolution.type: BY AUTHOR / Resolution: 26.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Hand-made, programs ) / Resolution.type: BY AUTHOR / Resolution: 26.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Hand-made, programs |
|---|
 Movie
Movie Controller
Controller