+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Local refinement of CIV from S. pombe | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Supercomplex / respiration / MEMBRANE PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Moe A / Brzezinski P | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structure and function of the III-IV-cyt supercomplex. Authors: Agnes Moe / Anna-Roza Dimogkioka / Doron Rapaport / Linda Näsvik Öjemyr / Peter Brzezinski /   Abstract: The respiratory chain in aerobic organisms is composed of a number of membrane-bound protein complexes that link electron transfer to proton translocation across the membrane. In mitochondria, the ...The respiratory chain in aerobic organisms is composed of a number of membrane-bound protein complexes that link electron transfer to proton translocation across the membrane. In mitochondria, the final electron acceptor, complex IV (CIV), receives electrons from dimeric complex III (CIII), via a mobile electron carrier, cytochrome . In the present study, we isolated the CIIICIV supercomplex from the fission yeast and determined its structure with bound cyt. using single-particle electron cryomicroscopy. A respiratory supercomplex factor 2 was found to be bound at CIV distally positioned in the supercomplex. In addition to the redox-active metal sites, we found a metal ion, presumably Zn, coordinated in the CIII subunit Cor1, which is encoded by the same gene () as the mitochondrial-processing peptidase subunit β. Our data show that the isolated CIIICIV supercomplex displays proteolytic activity suggesting a dual role of CIII in . As in the supercomplex from , subunit Cox5 of CIV faces towards one CIII monomer, but in the two complexes are rotated relative to each other by ~45°. This orientation yields equal distances between the cyt. binding sites at CIV and at each of the two CIII monomers. The structure shows cyt. bound at four positions, but only along one of the two symmetrical branches. Overall, this combined structural and functional study reveals the integration of peptidase activity with the CIII respiratory system and indicates a two-dimensional cyt. diffusion mechanism within the CIII-CIV supercomplex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18064.map.gz emd_18064.map.gz | 483 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18064-v30.xml emd-18064-v30.xml emd-18064.xml emd-18064.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
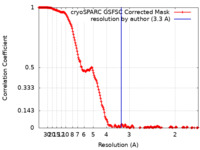
| FSC (resolution estimation) |  emd_18064_fsc.xml emd_18064_fsc.xml | 17.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_18064.png emd_18064.png | 47 KB | ||
| Filedesc metadata |  emd-18064.cif.gz emd-18064.cif.gz | 3.7 KB | ||
| Others |  emd_18064_half_map_1.map.gz emd_18064_half_map_1.map.gz emd_18064_half_map_2.map.gz emd_18064_half_map_2.map.gz | 474.4 MB 474.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18064 http://ftp.pdbj.org/pub/emdb/structures/EMD-18064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18064 | HTTPS FTP |
-Validation report
| Summary document |  emd_18064_validation.pdf.gz emd_18064_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18064_full_validation.pdf.gz emd_18064_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_18064_validation.xml.gz emd_18064_validation.xml.gz | 25.8 KB | Display | |
| Data in CIF |  emd_18064_validation.cif.gz emd_18064_validation.cif.gz | 33.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18064 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18064 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18064 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18064 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18064.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18064.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8464 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_18064_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
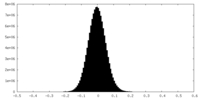
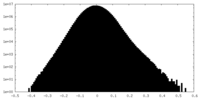
| Density Histograms |
-Half map: #2
| File | emd_18064_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
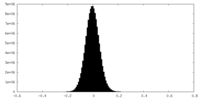
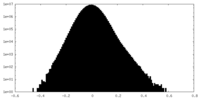
| Density Histograms |
- Sample components
Sample components
-Entire : III2-IV1 respiratory supercomplex
| Entire | Name: III2-IV1 respiratory supercomplex |
|---|---|
| Components |
|
-Supramolecule #1: III2-IV1 respiratory supercomplex
| Supramolecule | Name: III2-IV1 respiratory supercomplex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#23 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 41.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)