[English] 日本語
 Yorodumi
Yorodumi- EMDB-17865: Tilapia Lake Virus polymerase in vRNA elongation state with addit... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tilapia Lake Virus polymerase in vRNA elongation state with additional mode B promoter (transcriptase conformation) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral polymerase / VIRAL PROTEIN | |||||||||
| Function / homology | RNA-directed RNA polymerase activity / DNA/RNA polymerase superfamily / Uncharacterized protein / Putative PB1 / Uncharacterized protein Function and homology information Function and homology information | |||||||||
| Biological species |  Tilapia lake virus Tilapia lake virus | |||||||||
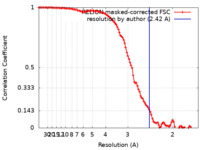
| Method | single particle reconstruction / cryo EM / Resolution: 2.42 Å | |||||||||
 Authors Authors | Arragain B / Cusack S | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural and functional analysis of the minimal orthomyxovirus-like polymerase of Tilapia Lake Virus from the highly diverged Amnoonviridae family. Authors: Benoit Arragain / Martin Pelosse / Albert Thompson / Stephen Cusack /   Abstract: Tilapia Lake Virus (TiLV), a recently discovered pathogen of tilapia fish, belongs to the Amnoonviridae family from the Articulavirales order. Its ten genome segments have characteristic conserved ...Tilapia Lake Virus (TiLV), a recently discovered pathogen of tilapia fish, belongs to the Amnoonviridae family from the Articulavirales order. Its ten genome segments have characteristic conserved ends and encode proteins with no known homologues, apart from the segment 1, which encodes an orthomyxo-like RNA-dependent-RNA polymerase core subunit. Here we show that segments 1-3 encode respectively the PB1, PB2 and PA-like subunits of an active heterotrimeric polymerase that maintains all domains found in the distantly related influenza polymerase, despite an unprecedented overall size reduction of 40%. Multiple high-resolution cryo-EM structures of TiLV polymerase in pre-initiation, initiation and active elongation states, show how it binds the vRNA and cRNA promoters and performs RNA synthesis, with both transcriptase and replicase configurations being characterised. However, the highly truncated endonuclease-like domain appears inactive and the putative cap-binding domain is autoinhibited, emphasising that many functional aspects of TiLV polymerase remain to be elucidated. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17865.map.gz emd_17865.map.gz | 9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17865-v30.xml emd-17865-v30.xml emd-17865.xml emd-17865.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17865_fsc.xml emd_17865_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_17865.png emd_17865.png | 87.1 KB | ||
| Masks |  emd_17865_msk_1.map emd_17865_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17865.cif.gz emd-17865.cif.gz | 7 KB | ||
| Others |  emd_17865_additional_1.map.gz emd_17865_additional_1.map.gz emd_17865_half_map_1.map.gz emd_17865_half_map_1.map.gz emd_17865_half_map_2.map.gz emd_17865_half_map_2.map.gz | 80.9 MB 81.1 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17865 http://ftp.pdbj.org/pub/emdb/structures/EMD-17865 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17865 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17865 | HTTPS FTP |
-Related structure data
| Related structure data |  8pszMC  8psnC  8psoC  8psqC  8pssC  8psuC  8psxC  8pt2C  8pt6C  8pt7C  8pthC  8ptjC  8qz8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17865.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17865.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17865_msk_1.map emd_17865_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_17865_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17865_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17865_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Tilapia lake virus polymerase
+Supramolecule #1: Tilapia lake virus polymerase
+Macromolecule #1: Polymerase acidic protein (PA-like)
+Macromolecule #2: Putative PB1
+Macromolecule #3: RNA-dependent RNA polymerase
+Macromolecule #4: 5' vRNA end - vRNA loop (40-mer)
+Macromolecule #5: Transcription-like product
+Macromolecule #6: ZINC ION
+Macromolecule #7: [(2~{R},3~{S},4~{R},5~{R})-5-(4-azanyl-2-oxidanylidene-pyrimidin-...
+Macromolecule #8: MAGNESIUM ION
+Macromolecule #9: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8psz: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)














































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)