[English] 日本語
 Yorodumi
Yorodumi- EMDB-17583: CHAPSO treated partial catalytic component (comprising only AnfD ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CHAPSO treated partial catalytic component (comprising only AnfD & AnfK, lacking AnfG and FeFeco) of iron nitrogenase from Rhodobacter capsulatus | |||||||||
 Map data Map data | Core complex of Fe-nitrogenase comprising only AnfD and AnfK. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nitrogen fixation / Fe nitrogenase / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationnitrogenase / nitrogenase activity / nitrogen fixation / iron-sulfur cluster binding / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) | |||||||||
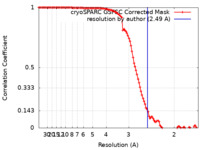
| Method | single particle reconstruction / cryo EM / Resolution: 2.49 Å | |||||||||
 Authors Authors | Schmidt FV / Schulz L / Zarzycki J / Prinz S / Erb TJ / Rebelein JG | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural insights into the iron nitrogenase complex. Authors: Frederik V Schmidt / Luca Schulz / Jan Zarzycki / Simone Prinz / Niels N Oehlmann / Tobias J Erb / Johannes G Rebelein /  Abstract: Nitrogenases are best known for catalyzing the reduction of dinitrogen to ammonia at a complex metallic cofactor. Recently, nitrogenases were shown to reduce carbon dioxide (CO) and carbon monoxide ...Nitrogenases are best known for catalyzing the reduction of dinitrogen to ammonia at a complex metallic cofactor. Recently, nitrogenases were shown to reduce carbon dioxide (CO) and carbon monoxide to hydrocarbons, offering a pathway to recycle carbon waste into hydrocarbon products. Among the three nitrogenase isozymes, the iron nitrogenase has the highest wild-type activity for the reduction of CO, but the molecular architecture facilitating these activities has remained unknown. Here, we report a 2.35-Å cryogenic electron microscopy structure of the ADP·AlF-stabilized iron nitrogenase complex from Rhodobacter capsulatus, revealing an [FeSC-(R)-homocitrate] cluster in the active site. The enzyme complex suggests that the iron nitrogenase G subunit is involved in cluster stabilization and substrate channeling and confers specificity between nitrogenase reductase and catalytic component proteins. Moreover, the structure highlights a different interface between the two catalytic halves of the iron and the molybdenum nitrogenase, potentially influencing the intrasubunit 'communication' and thus the nitrogenase mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17583.map.gz emd_17583.map.gz | 122.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17583-v30.xml emd-17583-v30.xml emd-17583.xml emd-17583.xml | 18.5 KB 18.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17583_fsc.xml emd_17583_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_17583.png emd_17583.png | 59.8 KB | ||
| Filedesc metadata |  emd-17583.cif.gz emd-17583.cif.gz | 6.3 KB | ||
| Others |  emd_17583_half_map_1.map.gz emd_17583_half_map_1.map.gz emd_17583_half_map_2.map.gz emd_17583_half_map_2.map.gz | 120.1 MB 120.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17583 http://ftp.pdbj.org/pub/emdb/structures/EMD-17583 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17583 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17583 | HTTPS FTP |
-Validation report
| Summary document |  emd_17583_validation.pdf.gz emd_17583_validation.pdf.gz | 891.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17583_full_validation.pdf.gz emd_17583_full_validation.pdf.gz | 891 KB | Display | |
| Data in XML |  emd_17583_validation.xml.gz emd_17583_validation.xml.gz | 19.3 KB | Display | |
| Data in CIF |  emd_17583_validation.cif.gz emd_17583_validation.cif.gz | 24.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17583 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17583 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17583 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17583 | HTTPS FTP |
-Related structure data
| Related structure data |  8pbbMC  8oieC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17583.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17583.map.gz / Format: CCP4 / Size: 129.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Core complex of Fe-nitrogenase comprising only AnfD and AnfK. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half-map A
| File | emd_17583_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map B
| File | emd_17583_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CHAPSO treated partial catalytic component (comprising only AnfD ...
| Entire | Name: CHAPSO treated partial catalytic component (comprising only AnfD & AnfK, lacking AnfG and FeFeco) of iron nitrogenase from Rhodobacter capsulatus |
|---|---|
| Components |
|
-Supramolecule #1: CHAPSO treated partial catalytic component (comprising only AnfD ...
| Supramolecule | Name: CHAPSO treated partial catalytic component (comprising only AnfD & AnfK, lacking AnfG and FeFeco) of iron nitrogenase from Rhodobacter capsulatus type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) |
-Macromolecule #1: Nitrogenase protein alpha chain
| Macromolecule | Name: Nitrogenase protein alpha chain / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) |
| Molecular weight | Theoretical: 60.221117 KDa |
| Recombinant expression | Organism:  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) |
| Sequence | String: MPYHEFEVSK CIPERREHAV MKAAGEDLTS CLPKGYLNTI PGTISERGCA YCGAKHVIGT PMKDVIHISH GPNGCTYDTW QTKRYISDN DNFQLKYTFA TDVKEKHVVF GAEGLLKKSM HEAFDAFPNI KRMTVYQTCT TALIGDDVDA IAKEVMEERG D VDVFVCNS ...String: MPYHEFEVSK CIPERREHAV MKAAGEDLTS CLPKGYLNTI PGTISERGCA YCGAKHVIGT PMKDVIHISH GPNGCTYDTW QTKRYISDN DNFQLKYTFA TDVKEKHVVF GAEGLLKKSM HEAFDAFPNI KRMTVYQTCT TALIGDDVDA IAKEVMEERG D VDVFVCNS PGFAGPSQSG GHHKINIAWL NQKVGTVEPD YLGEHVINYV GEYNIQGDQE VMIDYFNRMG IQVLSTFTGN GS YDSLRMM HRAHLNVLEC ARSAEYICDE LRARYGIPRL DIDGFGFEPL ANSLRKVALF FGIEDKAEAI IAEEYAKWKP QLD WYKERL KGKKVCLWPG GSKLWHWAHA IEEEMGLKVV SVYTKFGHQG DMEKGVSRCG EGALAIDDPN ELESVEAIEM LKPD IIFTG KRPGEFVKKH GVPYLNAHAY HNGPYKGFEG WVRFARDIYN AIYSPMRQLA ALDISAPDAA ITSGFRTAKM NADLT VSDE VKFSEVLHEY TGKYDSIAEI RARNQAYAAE QKALRDAVQP AAEWSHPQFE K UniProtKB: Nitrogenase protein alpha chain |
-Macromolecule #2: Nitrogenase iron-iron protein, beta subunit
| Macromolecule | Name: Nitrogenase iron-iron protein, beta subunit / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: nitrogenase |
|---|---|
| Source (natural) | Organism:  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) |
| Molecular weight | Theoretical: 50.776398 KDa |
| Recombinant expression | Organism:  Rhodobacter capsulatus SB 1003 (bacteria) Rhodobacter capsulatus SB 1003 (bacteria) |
| Sequence | String: MTCQVTQKAR EGTINPIFTC QPAGAQFASI GIKDCIGIVH GGQGCVMFVR LLISQHMKES FEIASSSVHE DGAVFGALDR VETAVEVLL TRYPDVKVVP IITTCSTEII GDDVDGLLSK LEDELLPTKF PGREVHLLTV HCPSFVGSMI TGYDKAVHDF V KKFATKDE ...String: MTCQVTQKAR EGTINPIFTC QPAGAQFASI GIKDCIGIVH GGQGCVMFVR LLISQHMKES FEIASSSVHE DGAVFGALDR VETAVEVLL TRYPDVKVVP IITTCSTEII GDDVDGLLSK LEDELLPTKF PGREVHLLTV HCPSFVGSMI TGYDKAVHDF V KKFATKDE PSDKINLITG WVNPGDVKEL KHLLEVMEVK ANVLFEVESF DSPLMPDLEH HSHGSTTIED LRDTANAKGT IA LNRYEGM KAADYLKKKF KVPAVIGPTP VGIRNTDAFL KAVSEMTGQP IPAQLVKERG LALDAIADIG HMFLADKRVA IYA NPDLAI GLTEFCLDLE MKPKLLLLGD DNSGYVKDPR VLALQENAPD LEIVTNADFW DLESRIQQGL ELDLILGHSK GRFI SIDYK VPMVRVGFPT YDRAGMYRHP VLGYGGAMFL AETMANTLFA DMEAKKNKEW ILNVW UniProtKB: Nitrogenase iron-iron protein, beta subunit |
-Macromolecule #3: FE(8)-S(7) CLUSTER
| Macromolecule | Name: FE(8)-S(7) CLUSTER / type: ligand / ID: 3 / Number of copies: 2 / Formula: CLF |
|---|---|
| Molecular weight | Theoretical: 671.215 Da |
| Chemical component information |  ChemComp-CLF: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL |
|---|---|
| Buffer | pH: 7.8 Details: 37.5 mM TRIS (pH = 7.8) 150 mM NaCl 3.75 mM sodium dithionite 0.4 % (m/V) CHAPSO |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 90 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)