+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Vaccinia Virus palisade layer A10 trimer | |||||||||
 Map data Map data | Final map after Phenix density modification of RELION-refined Vaccinia virus A10 trimer cryo-EM data. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Poxvirus / Vaccinia Virus / core / cryo-electron tomography / AlphaFold / VIRAL PROTEIN | |||||||||
| Function / homology | Poxvirus P4A / Poxvirus P4A protein / virion component / structural molecule activity / Major core protein OPG136 precursor Function and homology information Function and homology information | |||||||||
| Biological species |  Vaccinia virus Western Reserve Vaccinia virus Western Reserve | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Datler J / Hansen JM / Thader A / Schloegl A / Hodirnau VV / Schur FKM | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Multi-modal cryo-EM reveals trimers of protein A10 to form the palisade layer in poxvirus cores. Authors: Julia Datler / Jesse M Hansen / Andreas Thader / Alois Schlögl / Lukas W Bauer / Victor-Valentin Hodirnau / Florian K M Schur /  Abstract: Poxviruses are among the largest double-stranded DNA viruses, with members such as variola virus, monkeypox virus and the vaccination strain vaccinia virus (VACV). Knowledge about the structural ...Poxviruses are among the largest double-stranded DNA viruses, with members such as variola virus, monkeypox virus and the vaccination strain vaccinia virus (VACV). Knowledge about the structural proteins that form the viral core has remained sparse. While major core proteins have been annotated via indirect experimental evidence, their structures have remained elusive and they could not be assigned to individual core features. Hence, which proteins constitute which layers of the core, such as the palisade layer and the inner core wall, has remained enigmatic. Here we show, using a multi-modal cryo-electron microscopy (cryo-EM) approach in combination with AlphaFold molecular modeling, that trimers formed by the cleavage product of VACV protein A10 are the key component of the palisade layer. This allows us to place previously obtained descriptions of protein interactions within the core wall into perspective and to provide a detailed model of poxvirus core architecture. Importantly, we show that interactions within A10 trimers are likely generalizable over members of orthopox- and parapoxviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17410.map.gz emd_17410.map.gz | 5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17410-v30.xml emd-17410-v30.xml emd-17410.xml emd-17410.xml | 21.8 KB 21.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_17410.png emd_17410.png | 61.5 KB | ||
| Filedesc metadata |  emd-17410.cif.gz emd-17410.cif.gz | 6.5 KB | ||
| Others |  emd_17410_additional_1.map.gz emd_17410_additional_1.map.gz emd_17410_half_map_1.map.gz emd_17410_half_map_1.map.gz emd_17410_half_map_2.map.gz emd_17410_half_map_2.map.gz | 77.6 MB 99 MB 99.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17410 http://ftp.pdbj.org/pub/emdb/structures/EMD-17410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17410 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17410 | HTTPS FTP |
-Related structure data
| Related structure data |  8p4kMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17410.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17410.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map after Phenix density modification of RELION-refined Vaccinia virus A10 trimer cryo-EM data. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Final map after Phenix density modification of RELION-refined...
| File | emd_17410_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map after Phenix density modification of RELION-refined Vaccinia virus A10 trimer cryo-EM data. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Final map after Phenix density modification of RELION-refined...
| File | emd_17410_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map after Phenix density modification of RELION-refined Vaccinia virus A10 trimer cryo-EM data. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: 10A low-pass filtered map after Phenix density modification...
| File | emd_17410_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 10A low-pass filtered map after Phenix density modification of RELION-refined Vaccinia virus A10 trimer cryo-EM data. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Vaccinia virus Western Reserve
| Entire | Name:  Vaccinia virus Western Reserve Vaccinia virus Western Reserve |
|---|---|
| Components |
|
-Supramolecule #1: Vaccinia virus Western Reserve
| Supramolecule | Name: Vaccinia virus Western Reserve / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Purified from HeLa cells infected with Vaccinia Virus Western Reserve. NCBI-ID: 696871 / Sci species name: Vaccinia virus Western Reserve / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Core protein OPG136
| Macromolecule | Name: Core protein OPG136 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Vaccinia virus Western Reserve Vaccinia virus Western Reserve |
| Molecular weight | Theoretical: 69.068242 KDa |
| Sequence | String: MMPIKSIVTL DQLEDSEYLF RIVSTVLPHL CLDYKVCDQL KTTFVHPFDI LLNNSLGSVT KQDELQAAIS KLGINYLIDT TSRELKLFN VTLNAGNIDI INTPINISSE TNPIINTHSF YDLPPFTQHL LNIRLTDTEY RARFIGGYIK PDGSDSMDVL A EKKYPDLN ...String: MMPIKSIVTL DQLEDSEYLF RIVSTVLPHL CLDYKVCDQL KTTFVHPFDI LLNNSLGSVT KQDELQAAIS KLGINYLIDT TSRELKLFN VTLNAGNIDI INTPINISSE TNPIINTHSF YDLPPFTQHL LNIRLTDTEY RARFIGGYIK PDGSDSMDVL A EKKYPDLN FDNTYLFNIL YKDVINAPIK EFKAKIVNGV LSRQDFDNLI GVRQYITIQD RPRFDDAYNI ADAARHYGVN LN TLPLPNV DLTTMPTYKH LIMFEQYFIY TYDRVDIYYN GNKMLFDDEI INFTISMRYQ SLIPRLVDFF PDIPVNNNIV LHT RDPQNA AVNVTVALPN VQFVDINRNN KFFINFFNLL AKEQRSTAIK VTKSMFWDGM DYEEYKSKNL QDMMFINSTC YVFG LYNHN NTTYCSILSD IISAEKTPIR VCLLPRVVGG KTVTNLISET LKSISSMTIR EFPRKDKSIM HIGLSETGFM RFFQL LRLM ADKPHETAIK EVVMAYVGIK LGDKGSPYYI RKESYQDFIY LLFASMGFKV TTRRSIMGSN NISIISIRPR VTKQYI VAT LMKTSCSKNE AEKLITSAFD LLNFMVSVSD FRDYQS UniProtKB: Major core protein OPG136 precursor |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 9 Component:
Details: diluted 1:1 with 0.25% Trypsin (final concentration 0.125%) and 1:1 4% paraformaldehyde (final concentration 2%) in 1 mM Tris-HCl. | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 180 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 277 K / Instrument: LEICA EM GP / Details: Leica GP2. | |||||||||
| Details | Isolated viral cores with some detached soluble monodispersed particles. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number real images: 9264 / Average electron dose: 53.04 e/Å2 / Details: 34 frames total. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.25 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model / Details: version 2.3 |
|---|---|
| Details | Rosetta relaxation |
| Refinement | Protocol: OTHER |
| Output model |  PDB-8p4k: |
 Movie
Movie Controller
Controller







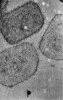

 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)