+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Rotavirus B NSP2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Octamer / RNA binding / Rotavirus / RNA chaperone / VIRAL PROTEIN | |||||||||
| Function / homology | Rotavirus non-structural protein 2 / hydrolase activity, acting on acid anhydrides / viral genome replication / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / host cell cytoplasm / RNA binding / ATP binding / metal ion binding / Non-structural protein 2 Function and homology information Function and homology information | |||||||||
| Biological species |  Rotavirus B / Rotavirus B /  Human rotavirus B strain CAL-1 Human rotavirus B strain CAL-1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Chamera S / Nowotny M | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J Virol / Year: 2024 Journal: J Virol / Year: 2024Title: Cryo-EM structure of rotavirus B NSP2 reveals its unique tertiary architecture. Authors: Sebastian Chamera / Krzysztof Wycisk / Mariusz Czarnocki-Cieciura / Marcin Nowotny /  Abstract: Rotavirus (RV) NSP2 is a multifunctional RNA chaperone that exhibits numerous activities that are essential for replication and viral genome packaging. We performed an analysis that highlighted a ...Rotavirus (RV) NSP2 is a multifunctional RNA chaperone that exhibits numerous activities that are essential for replication and viral genome packaging. We performed an analysis that highlighted a distant relationship of NSP2 from rotavirus B (RVB) to proteins from other human RVs. We solved a cryo-electron microscopy structure of RVB NSP2 that shows structural differences with corresponding proteins from other human RVs. Based on the structure, we identified amino acid residues that are involved in RNA interactions. Anisotropy titration experiments showed that these residues are important for nucleic acid binding. We also identified structural motifs that are conserved in all RV species. Collectively, our data complete the structural characterization of rotaviral NSP2 protein and demonstrate its structural diversity among RV species.IMPORTANCERotavirus B (RVB), also known as adult diarrhea rotavirus, has caused epidemics of severe diarrhea in China, India, and Bangladesh. Thousands of people are infected in a single RVB epidemic. However, information on this group of rotaviruses remains limited. As NSP2 is an essential protein in the viral life cycle, including its role in the formation of replication factories, it may be a target for future antiviral strategy against viruses with similar mechanisms. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17323.map.gz emd_17323.map.gz | 78.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17323-v30.xml emd-17323-v30.xml emd-17323.xml emd-17323.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_17323.png emd_17323.png | 49.9 KB | ||
| Filedesc metadata |  emd-17323.cif.gz emd-17323.cif.gz | 5.5 KB | ||
| Others |  emd_17323_half_map_1.map.gz emd_17323_half_map_1.map.gz emd_17323_half_map_2.map.gz emd_17323_half_map_2.map.gz | 77.6 MB 77.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17323 http://ftp.pdbj.org/pub/emdb/structures/EMD-17323 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17323 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17323 | HTTPS FTP |
-Related structure data
| Related structure data |  8p00MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17323.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17323.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17323_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
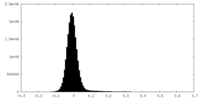
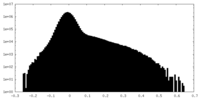
| Density Histograms |
-Half map: #1
| File | emd_17323_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
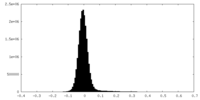
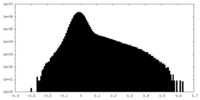
| Density Histograms |
- Sample components
Sample components
-Entire : Homooctamer of NSP2
| Entire | Name: Homooctamer of NSP2 |
|---|---|
| Components |
|
-Supramolecule #1: Homooctamer of NSP2
| Supramolecule | Name: Homooctamer of NSP2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Rotavirus B / Strain: Mexico Rotavirus B / Strain: Mexico |
-Macromolecule #1: Non-structural protein 2
| Macromolecule | Name: Non-structural protein 2 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Human rotavirus B strain CAL-1 Human rotavirus B strain CAL-1 |
| Molecular weight | Theoretical: 34.575672 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTQSVSLSDF IVKTEDGYMP SDRECVALDR YLSKEQKELR ETFKDGKNDR SALRIKMFLS PSPSRRFTQH GVVPMREIKT NTDIPSTLW TLVTDWLLNL LQDEENQEMF EDFISSKFPD VLASADKLAR FAQRLEDRKD VLHKNFSKAM NAFGACFWAI K PTFATEGK ...String: MTQSVSLSDF IVKTEDGYMP SDRECVALDR YLSKEQKELR ETFKDGKNDR SALRIKMFLS PSPSRRFTQH GVVPMREIKT NTDIPSTLW TLVTDWLLNL LQDEENQEMF EDFISSKFPD VLASADKLAR FAQRLEDRKD VLHKNFSKAM NAFGACFWAI K PTFATEGK CNVVRATDDS MILEFQPVPE YFRCGRSKAT FYKLYPLSDE QPVNGMLALK AVAGNQFFMY HGHGHIRTVP YH ELADAIK SYARKDKETL ESISKSPLAA QCGSKFLDML DGIRSKQKIE DVILKAKIFE KKRS UniProtKB: Non-structural protein 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.36 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: C-flat-2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.92 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 22131 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)