[English] 日本語
 Yorodumi
Yorodumi- EMDB-17286: Escherichia coli paused disome complex (rotated-PRE-1 | closed no... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Escherichia coli paused disome complex (rotated-PRE-1 | closed non-rotated PRE) | |||||||||
 Map data Map data | Cryo-EM reconstruction of the E. coli disome complex (rotated-PRE-1 | closed non-rotated PRE) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome / polysome / tranlsation / elongation / pausing / disome / collision / PURE system | |||||||||
| Biological species |  | |||||||||
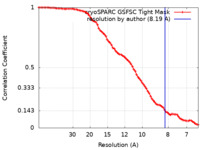
| Method | single particle reconstruction / cryo EM / Resolution: 8.19 Å | |||||||||
 Authors Authors | Fluegel T / Schacherl M | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM. Authors: Timo Flügel / Magdalena Schacherl / Anett Unbehaun / Birgit Schroeer / Marylena Dabrowski / Jörg Bürger / Thorsten Mielke / Thiemo Sprink / Christoph A Diebolder / Yollete V Guillén ...Authors: Timo Flügel / Magdalena Schacherl / Anett Unbehaun / Birgit Schroeer / Marylena Dabrowski / Jörg Bürger / Thorsten Mielke / Thiemo Sprink / Christoph A Diebolder / Yollete V Guillén Schlippe / Christian M T Spahn /  Abstract: Structural studies of translating ribosomes traditionally rely on in vitro assembly and stalling of ribosomes in defined states. To comprehensively visualize bacterial translation, we reactivated ex ...Structural studies of translating ribosomes traditionally rely on in vitro assembly and stalling of ribosomes in defined states. To comprehensively visualize bacterial translation, we reactivated ex vivo-derived E. coli polysomes in the PURE in vitro translation system and analyzed the actively elongating polysomes by cryo-EM. We find that 31% of 70S ribosomes assemble into disome complexes that represent eight distinct functional states including decoding and termination intermediates, and a pre-nucleophilic attack state. The functional diversity of disome complexes together with RNase digest experiments suggests that paused disome complexes transiently form during ongoing elongation. Structural analysis revealed five disome interfaces between leading and queueing ribosomes that undergo rearrangements as the leading ribosome traverses through the elongation cycle. Our findings reveal at the molecular level how bL9's CTD obstructs the factor binding site of queueing ribosomes to thwart harmful collisions and illustrate how translation dynamics reshape inter-ribosomal contacts. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17286.map.gz emd_17286.map.gz | 23.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17286-v30.xml emd-17286-v30.xml emd-17286.xml emd-17286.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17286_fsc.xml emd_17286_fsc.xml | 7.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_17286.png emd_17286.png | 79 KB | ||
| Masks |  emd_17286_msk_1.map emd_17286_msk_1.map | 48.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17286.cif.gz emd-17286.cif.gz | 4.4 KB | ||
| Others |  emd_17286_half_map_1.map.gz emd_17286_half_map_1.map.gz emd_17286_half_map_2.map.gz emd_17286_half_map_2.map.gz | 45.3 MB 45.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17286 http://ftp.pdbj.org/pub/emdb/structures/EMD-17286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17286 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17286.map.gz / Format: CCP4 / Size: 48.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17286.map.gz / Format: CCP4 / Size: 48.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of the E. coli disome complex (rotated-PRE-1 | closed non-rotated PRE) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.18 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17286_msk_1.map emd_17286_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM reconstruction of the E. coli disome complex...
| File | emd_17286_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of the E. coli disome complex (rotated-PRE-1 | closed non-rotated PRE). Half map A. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM reconstruction of the E. coli disome complex...
| File | emd_17286_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of the E. coli disome complex (rotated-PRE-1 | closed non-rotated PRE). Half map B. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia coli paused disome complex
| Entire | Name: Escherichia coli paused disome complex |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia coli paused disome complex
| Supramolecule | Name: Escherichia coli paused disome complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#60 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average exposure time: 1.13 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)