+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CAND1-CUL1-RBX1-SKP1-SKP2-DCNL1 | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | CAND1 / substrate receptor exchange factor / cullin-RING ligase / CRL / SCF / ligase / neddylation / DCNL1 co-E3 / ubiquitin signaling | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of protein neddylation / SCF complex assembly / ubiquitin-like protein binding / positive regulation of protein polyubiquitination / negative regulation of catalytic activity / Parkin-FBXW7-Cul1 ubiquitin ligase complex / cellular response to cell-matrix adhesion / F-box domain binding / Aberrant regulation of mitotic exit in cancer due to RB1 defects / regulation of protein neddylation ...positive regulation of protein neddylation / SCF complex assembly / ubiquitin-like protein binding / positive regulation of protein polyubiquitination / negative regulation of catalytic activity / Parkin-FBXW7-Cul1 ubiquitin ligase complex / cellular response to cell-matrix adhesion / F-box domain binding / Aberrant regulation of mitotic exit in cancer due to RB1 defects / regulation of protein neddylation / PcG protein complex / negative regulation of beige fat cell differentiation / cullin-RING-type E3 NEDD8 transferase / NEDD8 transferase activity / cullin-RING ubiquitin ligase complex / maintenance of protein location in nucleus / Cul7-RING ubiquitin ligase complex / cellular response to chemical stress / Loss of Function of FBXW7 in Cancer and NOTCH1 Signaling / positive regulation of protein autoubiquitination / RNA polymerase II transcription initiation surveillance / protein neddylation / ubiquitin conjugating enzyme binding / ubiquitin ligase activator activity / NEDD8 ligase activity / protein K27-linked ubiquitination / negative regulation of response to oxidative stress / VCB complex / Cul5-RING ubiquitin ligase complex / ubiquitin-ubiquitin ligase activity / ubiquitin-dependent protein catabolic process via the C-end degron rule pathway / positive regulation of intracellular estrogen receptor signaling pathway / SCF ubiquitin ligase complex / Cul2-RING ubiquitin ligase complex / Cul3-RING ubiquitin ligase complex / negative regulation of type I interferon production / SCF-dependent proteasomal ubiquitin-dependent protein catabolic process / Cul4A-RING E3 ubiquitin ligase complex / Cul4-RING E3 ubiquitin ligase complex / Prolactin receptor signaling / negative regulation of mitophagy / Cul4B-RING E3 ubiquitin ligase complex / ubiquitin ligase complex scaffold activity / regulation of protein ubiquitination / positive regulation of RNA polymerase II transcription preinitiation complex assembly / cullin family protein binding / protein monoubiquitination / ubiquitin ligase complex / protein K63-linked ubiquitination / positive regulation of double-strand break repair via homologous recombination / ubiquitin-like ligase-substrate adaptor activity / site of DNA damage / signal transduction in response to DNA damage / Nuclear events stimulated by ALK signaling in cancer / protein K48-linked ubiquitination / transcription-coupled nucleotide-excision repair / negative regulation of insulin receptor signaling pathway / regulation of cellular response to insulin stimulus / positive regulation of TORC1 signaling / post-translational protein modification / positive regulation of smooth muscle cell proliferation / intrinsic apoptotic signaling pathway / TBP-class protein binding / molecular function activator activity / animal organ morphogenesis / T cell activation / Regulation of BACH1 activity / MAP3K8 (TPL2)-dependent MAPK1/3 activation / negative regulation of canonical NF-kappaB signal transduction / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / cellular response to amino acid stimulus / Vpu mediated degradation of CD4 / Dectin-1 mediated noncanonical NF-kB signaling / Degradation of DVL / Degradation of CRY and PER proteins / G1/S transition of mitotic cell cycle / negative regulation of canonical Wnt signaling pathway / Activation of NF-kappaB in B cells / Iron uptake and transport / Degradation of GLI1 by the proteasome / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Recognition of DNA damage by PCNA-containing replication complex / Negative regulation of NOTCH4 signaling / RING-type E3 ubiquitin transferase / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Hedgehog 'on' state / Vif-mediated degradation of APOBEC3G / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / beta-catenin binding / NOTCH1 Intracellular Domain Regulates Transcription / Degradation of beta-catenin by the destruction complex / Evasion by RSV of host interferon responses / DNA Damage Recognition in GG-NER / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / G2/M transition of mitotic cell cycle / CLEC7A (Dectin-1) signaling Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
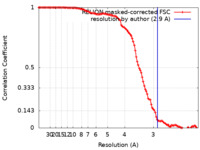
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||
 Authors Authors | Shaaban M / Clapperton JA / Ding S / Maeots ME / Enchev RI / Maslen S / Skehel JM | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2023 Journal: Mol Cell / Year: 2023Title: Structural and mechanistic insights into the CAND1-mediated SCF substrate receptor exchange. Authors: Mohammed Shaaban / Julie A Clapperton / Shan Ding / Simone Kunzelmann / Märt-Erik Mäeots / Sarah L Maslen / J Mark Skehel / Radoslav I Enchev /  Abstract: Modular SCF (SKP1-CUL1-Fbox) ubiquitin E3 ligases orchestrate multiple cellular pathways in eukaryotes. Their variable SKP1-Fbox substrate receptor (SR) modules enable regulated substrate recruitment ...Modular SCF (SKP1-CUL1-Fbox) ubiquitin E3 ligases orchestrate multiple cellular pathways in eukaryotes. Their variable SKP1-Fbox substrate receptor (SR) modules enable regulated substrate recruitment and subsequent proteasomal degradation. CAND proteins are essential for the efficient and timely exchange of SRs. To gain structural understanding of the underlying molecular mechanism, we reconstituted a human CAND1-driven exchange reaction of substrate-bound SCF alongside its co-E3 ligase DCNL1 and visualized it by cryo-EM. We describe high-resolution structural intermediates, including a ternary CAND1-SCF complex, as well as conformational and compositional intermediates representing SR- or CAND1-dissociation. We describe in molecular detail how CAND1-induced conformational changes in CUL1/RBX1 provide an optimized DCNL1-binding site and reveal an unexpected dual role for DCNL1 in CAND1-SCF dynamics. Moreover, a partially dissociated CAND1-SCF conformation accommodates cullin neddylation, leading to CAND1 displacement. Our structural findings, together with functional biochemical assays, help formulate a detailed model for CAND-SCF regulation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17116.map.gz emd_17116.map.gz | 110.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17116-v30.xml emd-17116-v30.xml emd-17116.xml emd-17116.xml | 27.1 KB 27.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17116_fsc.xml emd_17116_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_17116.png emd_17116.png | 123.8 KB | ||
| Filedesc metadata |  emd-17116.cif.gz emd-17116.cif.gz | 8.7 KB | ||
| Others |  emd_17116_half_map_1.map.gz emd_17116_half_map_1.map.gz emd_17116_half_map_2.map.gz emd_17116_half_map_2.map.gz | 98.3 MB 98.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17116 http://ftp.pdbj.org/pub/emdb/structures/EMD-17116 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17116 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17116 | HTTPS FTP |
-Related structure data
| Related structure data |  8or3MC  8or0C  8or2C  8or4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17116.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17116.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17116_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17116_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CAND1-CUL1-RBX1-SKP1-SKP2-CKS1-CDK2-CyclinE-DCNL1
| Entire | Name: CAND1-CUL1-RBX1-SKP1-SKP2-CKS1-CDK2-CyclinE-DCNL1 |
|---|---|
| Components |
|
-Supramolecule #1: CAND1-CUL1-RBX1-SKP1-SKP2-CKS1-CDK2-CyclinE-DCNL1
| Supramolecule | Name: CAND1-CUL1-RBX1-SKP1-SKP2-CKS1-CDK2-CyclinE-DCNL1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Cullin-1
| Macromolecule | Name: Cullin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 93.730672 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MWSHPQFEKG SAGSAAGSGA GWSHPQFEKL EVLFQGPMSS TRSQNPHGLK QIGLDQIWDD LRAGIQQVYT RQSMAKSRYM ELYTHVYNY CTSVHQSNQA RGAGVPPSKS KKGQTPGGAQ FVGLELYKRL KEFLKNYLTN LLKDGEDLMD ESVLKFYTQQ W EDYRFSSK ...String: MWSHPQFEKG SAGSAAGSGA GWSHPQFEKL EVLFQGPMSS TRSQNPHGLK QIGLDQIWDD LRAGIQQVYT RQSMAKSRYM ELYTHVYNY CTSVHQSNQA RGAGVPPSKS KKGQTPGGAQ FVGLELYKRL KEFLKNYLTN LLKDGEDLMD ESVLKFYTQQ W EDYRFSSK VLNGICAYLN RHWVRRECDE GRKGIYEIYS LALVTWRDCL FRPLNKQVTN AVLKLIEKER NGETINTRLI SG VVQSYVE LGLNEDDAFA KGPTLTVYKE SFESQFLADT ERFYTRESTE FLQQNPVTEY MKKAEARLLE EQRRVQVYLH EST QDELAR KCEQVLIEKH LEIFHTEFQN LLDADKNEDL GRMYNLVSRI QDGLGELKKL LETHIHNQGL AAIEKCGEAA LNDP KMYVQ TVLDVHKKYN ALVMSAFNND AGFVAALDKA CGRFINNNAV TKMAQSSSKS PELLARYCDS LLKKSSKNPE EAELE DTLN QVMVVFKYIE DKDVFQKFYA KMLAKRLVHQ NSASDDAEAS MISKLKQACG FEYTSKLQRM FQDIGVSKDL NEQFKK HLT NSEPLDLDFS IQVLSSGSWP FQQSCTFALP SELERSYQRF TAFYASRHSG RKLTWLYQLS KGELVTNCFK NRYTLQA ST FQMAILLQYN TEDAYTVQQL TDSTQIKMDI LAQVLQILLK SKLLVLEDEN ANVDEVELKP DTLIKLYLGY KNKKLRVN I NVPMKTEQKQ EQETTHKNIE EDRKLLIQAA IVRIMKMRKV LKHQQLLGEV LTQLSSRFKP RVPVIKKCID ILIEKEYLE RVDGEKDTYS YLA UniProtKB: Cullin-1 |
-Macromolecule #2: E3 ubiquitin-protein ligase RBX1
| Macromolecule | Name: E3 ubiquitin-protein ligase RBX1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.289977 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAAMDVDTP SGTNSGAGKK RFEVKKWNAV ALWAWDIVVD NCAICRNHIM DLCIECQANQ ASATSEECTV AWGVCNHAFH FHCISRWLK TRQVCPLDNR EWEFQKYGH UniProtKB: E3 ubiquitin-protein ligase RBX1 |
-Macromolecule #3: Cullin-associated NEDD8-dissociated protein 1
| Macromolecule | Name: Cullin-associated NEDD8-dissociated protein 1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 137.358219 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSPEFPGRMA SASYHISNLL EKMTSSDKDF RFMATNDLMT ELQKDSIKLD DDSERKVVKM ILKLLEDKNG EVQNLAVKCL GPLVSKVKE YQVETIVDTL CTNMLSDKEQ LRDISSIGLK TVIGELPPAS SGSALAANVC KKITGRLTSA IAKQEDVSVQ L EALDIMAD ...String: GSPEFPGRMA SASYHISNLL EKMTSSDKDF RFMATNDLMT ELQKDSIKLD DDSERKVVKM ILKLLEDKNG EVQNLAVKCL GPLVSKVKE YQVETIVDTL CTNMLSDKEQ LRDISSIGLK TVIGELPPAS SGSALAANVC KKITGRLTSA IAKQEDVSVQ L EALDIMAD MLSRQGGLLV NFHPSILTCL LPQLTSPRLA VRKRTIIALG HLVMSCGNIV FVDLIEHLLS ELSKNDSMST TR TYIQCIA AISRQAGHRI GEYLEKIIPL VVKFCNVDDD ELREYCIQAF ESFVRRCPKE VYPHVSTIIN ICLKYLTYDP NYN YDDEDE DENAMDADGG DDDDQGSDDE YSDDDDMSWK VRRAAAKCLD AVVSTRHEML PEFYKTVSPA LISRFKEREE NVKA DVFHA YLSLLKQTRP VQSWLCDPDA MEQGETPLTM LQSQVPNIVK ALHKQMKEKS VKTRQCCFNM LTELVNVLPG ALTQH IPVL VPGIIFSLND KSSSSNLKID ALSCLYVILC NHSPQVFHPH VQALVPPVVA CVGDPFYKIT SEALLVTQQL VKVIRP LDQ PSSFDATPYI KDLFTCTIKR LKAADIDQEV KERAISCMGQ IICNLGDNLG SDLPNTLQIF LERLKNEITR LTTVKAL TL IAGSPLKIDL RPVLGEGVPI LASFLRKNQR ALKLGTLSAL DILIKNYSDS LTAAMIDAVL DELPPLISES DMHVSQMA I SFLTTLAKVY PSSLSKISGS ILNELIGLVR SPLLQGGALS AMLDFFQALV VTGTNNLGYM DLLRMLTGPV YSQSTALTH KQSYYSIAKC VAALTRACPK EGPAVVGQFI QDVKNSRSTD SIRLLALLSL GEVGHHIDLS GQLELKSVIL EAFSSPSEEV KSAASYALG SISVGNLPEY LPFVLQEITS QPKRQYLLLH SLKEIISSAS VVGLKPYVEN IWALLLKHCE CAEEGTRNVV A ECLGKLTL IDPETLLPRL KGYLISGSSY ARSSVVTAVK FTISDHPQPI DPLLKNCIGD FLKTLEDPDL NVRRVALVTF NS AAHNKPS LIRDLLDTVL PHLYNETKVR KELIREVEMG PFKHTVDDGL DIRKAAFECM YTLLDSCLDR LDIFEFLNHV EDG LKDHYD IKMLTFLMLV RLSTLCPSAV LQRLDRLVEP LRATCTTKVK ANSVKQEFEK QDELKRSAMR AVAALLTIPE AEKS PLMSE FQSQISSNPE LAAIFESIQK DSSSTNLESM DTS UniProtKB: Cullin-associated NEDD8-dissociated protein 1 |
-Macromolecule #4: S-phase kinase-associated protein 1
| Macromolecule | Name: S-phase kinase-associated protein 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.679965 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPSIKLQSSD GEIFEVDVEI AKQSVTIKTM LEDLGMDDEG DDDPVPLPNV NAAILKKVIQ WCTHHKDDPP PPEDDENKEK RTDDIPVWD QEFLKVDQGT LFELILAANY LDIKGLLDVT CKTVANMIKG KTPEEIRKTF NIKNDFTEEE EAQVRKENQW C EEK UniProtKB: S-phase kinase-associated protein 1 |
-Macromolecule #5: S-phase kinase-associated protein 2
| Macromolecule | Name: S-phase kinase-associated protein 2 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 48.335312 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSPEFMHRKH LQEIPDLSSN VATSFTWGWD SSKTSELLSG MGVSALEKEE PDSENIPQEL LSNLGHPESP PRKRLKSKGS DKDFVIVRR PKLNRENFPG VSWDSLPDEL LLGIFSCLCL PELLKVSGVC KRWYRLASDE SLWQTLDLTG KNLHPDVTGR L LSQGVIAF ...String: GSPEFMHRKH LQEIPDLSSN VATSFTWGWD SSKTSELLSG MGVSALEKEE PDSENIPQEL LSNLGHPESP PRKRLKSKGS DKDFVIVRR PKLNRENFPG VSWDSLPDEL LLGIFSCLCL PELLKVSGVC KRWYRLASDE SLWQTLDLTG KNLHPDVTGR L LSQGVIAF RCPRSFMDQP LAEHFSPFRV QHMDLSNSVI EVSTLHGILS QCSKLQNLSL EGLRLSDPIV NTLAKNSNLV RL NLSGCSG FSEFALQTLL SSCSRLDELN LSWCFDFTEK HVQVAVAHVS ETITQLNLSG YRKNLQKSDL STLVRRCPNL VHL DLSDSV MLKNDCFQEF FQLNYLQHLS LSRCYDIIPE TLLELGEIPT LKTLQVFGIV PDGTLQLLKE ALPHLQINCS HFTT IARPT IGNKKNQEIW GIKCRLTLQK PSCL UniProtKB: S-phase kinase-associated protein 2 |
-Macromolecule #6: DCN1-like protein 1
| Macromolecule | Name: DCN1-like protein 1 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 30.304439 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSMNKLKSSQ KDKVRQFMIF TQSSEKTAVS CLSQNDWKLD VATDNFFQNP ELYIRESVKG SLDRKKLEQL YNRYKDPQDE NKIGIDGIQ QFCDDLALDP ASISVLIIAW KFRAATQCEF SKQEFMDGMT ELGCDSIEKL KAQIPKMEQE LKEPGRFKDF Y QFTFNFAK ...String: GSMNKLKSSQ KDKVRQFMIF TQSSEKTAVS CLSQNDWKLD VATDNFFQNP ELYIRESVKG SLDRKKLEQL YNRYKDPQDE NKIGIDGIQ QFCDDLALDP ASISVLIIAW KFRAATQCEF SKQEFMDGMT ELGCDSIEKL KAQIPKMEQE LKEPGRFKDF Y QFTFNFAK NPGQKGLDLE MAIAYWNLVL NGRFKFLDLW NKFLLEHHKR SIPKDTWNLL LDFSTMIADD MSNYDEEGAW PV LIDDFVE FARPQIAGTK STTV UniProtKB: DCN1-like protein 1 |
-Macromolecule #7: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 7 / Number of copies: 3 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 49.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 130000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)