+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | virus enhancing amyloid fibril formed by CKFKFQF | |||||||||
 Map data Map data | postprocessed map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | virus enhancing amyloid fibril / protein fibril / prion | |||||||||
| Biological species | synthetic construct (others) / HIV whole-genome vector AA1305#18 (others) | |||||||||
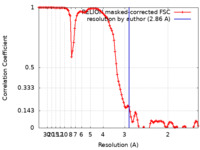
| Method | helical reconstruction / cryo EM / Resolution: 2.86 Å | |||||||||
 Authors Authors | Heerde T / Schmidt M / Faendrich M | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure and polymorphic maturation of a viral transduction enhancing amyloid fibril. Authors: Thomas Heerde / Desiree Schütz / Yu-Jie Lin / Jan Münch / Matthias Schmidt / Marcus Fändrich /  Abstract: Amyloid fibrils have emerged as innovative tools to enhance the transduction efficiency of retroviral vectors in gene therapy strategies. In this study, we used cryo-electron microscopy to analyze ...Amyloid fibrils have emerged as innovative tools to enhance the transduction efficiency of retroviral vectors in gene therapy strategies. In this study, we used cryo-electron microscopy to analyze the structure of a biotechnologically engineered peptide fibril that enhances retroviral infectivity. Our findings show that the peptide undergoes a time-dependent morphological maturation into polymorphic amyloid fibril structures. The fibrils consist of mated cross-β sheets that interact by the hydrophobic residues of the amphipathic fibril-forming peptide. The now available structural data help to explain the mechanism of retroviral infectivity enhancement, provide insights into the molecular plasticity of amyloid structures and illuminate the thermodynamic basis of their morphological maturation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16930.map.gz emd_16930.map.gz | 4.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16930-v30.xml emd-16930-v30.xml emd-16930.xml emd-16930.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16930_fsc.xml emd_16930_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16930.png emd_16930.png | 53.9 KB | ||
| Masks |  emd_16930_msk_1.map emd_16930_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16930.cif.gz emd-16930.cif.gz | 5 KB | ||
| Others |  emd_16930_additional_1.map.gz emd_16930_additional_1.map.gz emd_16930_half_map_1.map.gz emd_16930_half_map_1.map.gz emd_16930_half_map_2.map.gz emd_16930_half_map_2.map.gz | 95.2 MB 79.2 MB 79.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16930 http://ftp.pdbj.org/pub/emdb/structures/EMD-16930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16930 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16930.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16930.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocessed map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.81 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16930_msk_1.map emd_16930_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
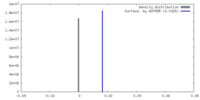
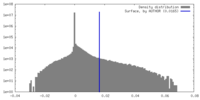
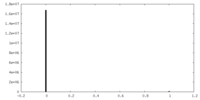
| Density Histograms |
-Additional map: Unmasked map
| File | emd_16930_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unmasked map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: second half-map
| File | emd_16930_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | second half-map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: first half-map
| File | emd_16930_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | first half-map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : virus enhancing amyloid
| Entire | Name: virus enhancing amyloid |
|---|---|
| Components |
|
-Supramolecule #1: virus enhancing amyloid
| Supramolecule | Name: virus enhancing amyloid / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: PNF-18
| Macromolecule | Name: PNF-18 / type: protein_or_peptide / ID: 1 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: HIV whole-genome vector AA1305#18 (others) |
| Molecular weight | Theoretical: 949.168 Da |
| Sequence | String: CKFKFQF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7 / Component - Concentration: 50.0 mM / Component - Formula: C8H18N2O4S Component - Name: 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid Details: 50 mM 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid |
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 96 % / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 8.0 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: BACKBONE TRACE / Target criteria: correlation coefficient |
|---|---|
| Output model |  PDB-8okr: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)