+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ivabradine bound to HCN4 channel | |||||||||||||||||||||
 Map data Map data | sharped map | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | HCN channels / ivabradine / ion transport / MEMBRANE PROTEIN / drug / hearth / hcn4 | |||||||||||||||||||||
| Biological species |  | |||||||||||||||||||||
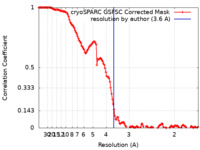
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||||||||||||||
 Authors Authors | Saponaro A / Chaves-Sanjuan A / Sharifzadeh AS / Clarke OB / Marabelli C / Bolognesi M / Thiel G / Moroni A | |||||||||||||||||||||
| Funding support |  France, France,  Italy, European Union, Italy, European Union,  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2024 Journal: Proc Natl Acad Sci U S A / Year: 2024Title: Structural determinants of ivabradine block of the open pore of HCN4. Authors: Andrea Saponaro / Jan H Krumbach / Antonio Chaves-Sanjuan / Atiyeh Sadat Sharifzadeh / Alessandro Porro / Roberta Castelli / Kay Hamacher / Martino Bolognesi / Dario DiFrancesco / Oliver B ...Authors: Andrea Saponaro / Jan H Krumbach / Antonio Chaves-Sanjuan / Atiyeh Sadat Sharifzadeh / Alessandro Porro / Roberta Castelli / Kay Hamacher / Martino Bolognesi / Dario DiFrancesco / Oliver B Clarke / Gerhard Thiel / Anna Moroni /    Abstract: HCN1-4 channels are the molecular determinants of the I/I current that crucially regulates cardiac and neuronal cell excitability. HCN dysfunctions lead to sinoatrial block (HCN4), epilepsy (HCN1), ...HCN1-4 channels are the molecular determinants of the I/I current that crucially regulates cardiac and neuronal cell excitability. HCN dysfunctions lead to sinoatrial block (HCN4), epilepsy (HCN1), and chronic pain (HCN2), widespread medical conditions awaiting subtype-specific treatments. Here, we address the problem by solving the cryo-EM structure of HCN4 in complex with ivabradine, to date the only HCN-specific drug on the market. Our data show ivabradine bound inside the open pore at 3 Å resolution. The structure unambiguously proves that Y507 and I511 on S6 are the molecular determinants of ivabradine binding to the inner cavity, while F510, pointing outside the pore, indirectly contributes to the block by controlling Y507. Cysteine 479, unique to the HCN selectivity filter (SF), accelerates the kinetics of block. Molecular dynamics simulations further reveal that ivabradine blocks the permeating ion inside the SF by electrostatic repulsion, a mechanism previously proposed for quaternary ammonium ions. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16860.map.gz emd_16860.map.gz | 230.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16860-v30.xml emd-16860-v30.xml emd-16860.xml emd-16860.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16860_fsc.xml emd_16860_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_16860.png emd_16860.png | 170.8 KB | ||
| Masks |  emd_16860_msk_1.map emd_16860_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16860.cif.gz emd-16860.cif.gz | 7 KB | ||
| Others |  emd_16860_additional_1.map.gz emd_16860_additional_1.map.gz emd_16860_half_map_1.map.gz emd_16860_half_map_1.map.gz emd_16860_half_map_2.map.gz emd_16860_half_map_2.map.gz | 120.3 MB 226.6 MB 226.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16860 http://ftp.pdbj.org/pub/emdb/structures/EMD-16860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16860 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16860.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16860.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharped map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
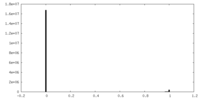
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16860_msk_1.map emd_16860_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharped map
| File | emd_16860_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharped map | ||||||||||||
| Projections & Slices |
| ||||||||||||
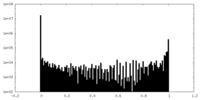
| Density Histograms |
-Half map: half a
| File | emd_16860_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half a | ||||||||||||
| Projections & Slices |
| ||||||||||||
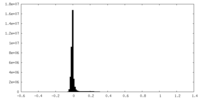
| Density Histograms |
-Half map: half b
| File | emd_16860_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half b | ||||||||||||
| Projections & Slices |
| ||||||||||||
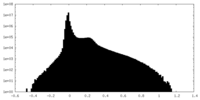
| Density Histograms |
- Sample components
Sample components
-Entire : HCN4
| Entire | Name: HCN4 |
|---|---|
| Components |
|
-Supramolecule #1: HCN4
| Supramolecule | Name: HCN4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: HCN4 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 400 KDa |
-Macromolecule #1: Potassium/sodium hyperpolarization-activated cyclic nucleotide-ga...
| Macromolecule | Name: Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 98.522727 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDKLPPSMRK RLYSLPQQVG AKAWIMDEEE DAEEEGAGGR QDPRRRSIRL RPLPSPSPSP SAAAAAAGGA ESRGAALGGA ADGEGPARG AAKSSTNGDC RRFRGSLASL GSRGGGGGGG STGGGSHGHL HDSAEERRLI AEGDASPGED RTPPGLAAEP E RPGAPAPP ...String: MDKLPPSMRK RLYSLPQQVG AKAWIMDEEE DAEEEGAGGR QDPRRRSIRL RPLPSPSPSP SAAAAAAGGA ESRGAALGGA ADGEGPARG AAKSSTNGDC RRFRGSLASL GSRGGGGGGG STGGGSHGHL HDSAEERRLI AEGDASPGED RTPPGLAAEP E RPGAPAPP AASPPQVPSS CGEQRPADAA VKVEGGAAAG DQILPEAEAR LGQAGFMQRQ FGAMLQPGVN KFSLRMFGSQ KA VEREQER VKSAGFWIIH PYSDFRFYWD LTMLLLMVGN LIIIPVGITF FKDENTTPWI VFNVVSDTFF LIDLVLNFRT GIV VEDNTD IILDPRRIKM KYLKSWFVVD FVSSIPVDYI FLIVETRIDS EVYKTARALR IVRFTKILSL LRLLRLSRLI RYIH QWEEI FHMTYDLASA VVRIVNLIGM MLLLCHWDGC LQFLVPMLQD FPDDCWVSLN NMVNNSWGKQ YSYALFKAMS HMLCI GYGR QAPMGMSDVW LTMLSMIVGA TCYAMFIGHA TALIQSLDSS RRQYQEKYKQ VEQYMSFHKL PPDTRQRIHD YYEHRY QGK MFDEESILGE LSEPLREEII NFNCRKLVAS MPLFANADPN FVTSMLTKLR FEVFQPGDYI IREGTIGKKM YFIQHGV VS VLTKGNKETK LADGSYFGEI CLLTRGRRTA SVRADTYCRL YSLSVDNFNE VLEEYPMMRR AFETVALDRL DRIGKKNS I HKVQHDLSSG VSNYQENAIV QRIVQHDREM AHCARRAQAT TPVAPAIWTP LIQAPLQAAA QDLKLISASQ PALPQDGAQ TLRRASPHSS SGESVAALPP AGGPFPRAPG RPPGAGPGQH VTLTLPRKAS SGSLPPPLSL FGPRAAPRLT AAPQREPGAK SEPVRSKLP SNL |
-Macromolecule #2: Ivabradine
| Macromolecule | Name: Ivabradine / type: ligand / ID: 2 / Number of copies: 1 / Formula: VNZ |
|---|---|
| Molecular weight | Theoretical: 468.585 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7 / Component - Concentration: 200.0 mM / Component - Formula: NaCl / Component - Name: sodium chloride |
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: 30mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 302 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 6000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number grids imaged: 1 / Number real images: 31670 / Average exposure time: 1.7 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)