+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 21S ribosomal precursors induced by heat shock. | |||||||||
 Map data Map data | 21S cryo-EM map, postprocessed and masked | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | 21S / DnaK / Heat shock / Ribosome assembly / RIBOSOME | |||||||||
| Biological species |  | |||||||||
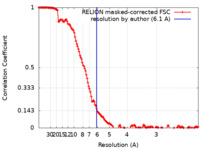
| Method | single particle reconstruction / cryo EM / Resolution: 6.1 Å | |||||||||
 Authors Authors | Giudice E / Georgeault S / Lavigne R / Pineau C / Trautwetter A / Ermel G / Blanco C / Gillet R | |||||||||
| Funding support |  France, 2 items France, 2 items
| |||||||||
 Citation Citation |  Journal: Int J Mol Sci / Year: 2023 Journal: Int J Mol Sci / Year: 2023Title: Purification and Characterization of Authentic 30S Ribosomal Precursors Induced by Heat Shock. Authors: Emmanuel Giudice / Sylvie Georgeault / Régis Lavigne / Charles Pineau / Annie Trautwetter / Gwennola Ermel / Carlos Blanco / Reynald Gillet /  Abstract: Ribosome biogenesis is a complex and multistep process that depends on various assembly factors. To understand this process and identify the ribosome assembly intermediates, most studies have set out ...Ribosome biogenesis is a complex and multistep process that depends on various assembly factors. To understand this process and identify the ribosome assembly intermediates, most studies have set out to delete or deplete these assembly factors. Instead, we took advantage of the impact of heat stress (45 °C) on the late stages of the biogenesis of the 30S ribosomal subunit to explore authentic precursors. Under these conditions, reduced levels of the DnaK chaperone proteins devoted to ribosome assembly lead to the transient accumulation of 21S ribosomal particles, which are 30S precursors. We constructed strains with different affinity tags on one early and one late 30S ribosomal protein and purified the 21S particles that form under heat shock. A combination of relative quantification using mass spectrometry-based proteomics and cryo-electron microscopy (cryo-EM) was then used to determine their protein contents and structures. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16556.map.gz emd_16556.map.gz | 17.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16556-v30.xml emd-16556-v30.xml emd-16556.xml emd-16556.xml | 37.5 KB 37.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16556_fsc.xml emd_16556_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_16556.png emd_16556.png | 61.4 KB | ||
| Masks |  emd_16556_msk_1.map emd_16556_msk_1.map emd_16556_msk_2.map emd_16556_msk_2.map emd_16556_msk_3.map emd_16556_msk_3.map | 244.1 MB 244.1 MB 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16556.cif.gz emd-16556.cif.gz | 5.5 KB | ||
| Others |  emd_16556_additional_1.map.gz emd_16556_additional_1.map.gz emd_16556_additional_2.map.gz emd_16556_additional_2.map.gz emd_16556_additional_3.map.gz emd_16556_additional_3.map.gz emd_16556_additional_4.map.gz emd_16556_additional_4.map.gz emd_16556_additional_5.map.gz emd_16556_additional_5.map.gz emd_16556_additional_6.map.gz emd_16556_additional_6.map.gz emd_16556_additional_7.map.gz emd_16556_additional_7.map.gz emd_16556_additional_8.map.gz emd_16556_additional_8.map.gz emd_16556_additional_9.map.gz emd_16556_additional_9.map.gz emd_16556_half_map_1.map.gz emd_16556_half_map_1.map.gz emd_16556_half_map_2.map.gz emd_16556_half_map_2.map.gz | 193.7 MB 183.6 MB 14.9 MB 183 MB 183.7 MB 161.5 MB 11.9 MB 161.5 MB 161.5 MB 194.3 MB 194.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16556 http://ftp.pdbj.org/pub/emdb/structures/EMD-16556 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16556 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16556 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16556.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16556.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 21S cryo-EM map, postprocessed and masked | ||||||||||||||||||||||||||||||||||||
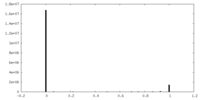
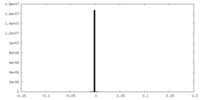
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Mask #1
+Mask #2
+Mask #3
+Additional map: 21S cryo-EM map (not postprocessed)
+Additional map: multibody analysis : 21S Body first half map
+Additional map: multibody analysis : 21S Body, postprocessed and masked
+Additional map: multibody analysis : 21S Body (not postprocessed)
+Additional map: multibody analysis : 21S Body second half map
+Additional map: multibody analysis : 21S Head (not postprocessed)
+Additional map: multibody analysis : 21S Head, postprocessed and masked
+Additional map: multibody analysis : 21S Head first half map
+Additional map: multibody analysis : 21S Head second half map
+Half map: 21S first half map
+Half map: 21S second half map
- Sample components
Sample components
-Entire : 21S ribosomal precursors induced by heat shock.
| Entire | Name: 21S ribosomal precursors induced by heat shock. |
|---|---|
| Components |
|
-Supramolecule #1: 21S ribosomal precursors induced by heat shock.
| Supramolecule | Name: 21S ribosomal precursors induced by heat shock. / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: 16S ribosomal RNA
| Supramolecule | Name: 16S ribosomal RNA / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: 21S Ribosomal proteins
| Supramolecule | Name: 21S Ribosomal proteins / type: complex / ID: 3 / Parent: 1 Details: uS3 uS4 uS5 bS6 uS7 uS8 uS9 uS10 uS11 uS12 uS13 uS14 uS15 bS16 uS17 bS18 uS19 bS20 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.04 kPa / Details: 30mA | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295.15 K / Instrument: FEI VITROBOT MARK III / Details: blot for 2s before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Microscope was modified with a Cs corrector |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 922 / Average exposure time: 1.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 2.7 µm / Calibrated defocus min: 0.4 µm / Calibrated magnification: 120443 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.0023 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)