[English] 日本語
 Yorodumi
Yorodumi- EMDB-16534: human alpha7 nicotinic receptor in complex with the C4 nanobody a... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | human alpha7 nicotinic receptor in complex with the C4 nanobody and nicotine | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | ion channel nanobody Nicotinic acetylcholine receptor / MEMBRANE PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationsensory processing / synaptic transmission involved in micturition / response to acetylcholine / dendrite arborization / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / acetylcholine receptor activity / acetylcholine-gated channel complex / chloride channel regulator activity / regulation of amyloid fibril formation / acetylcholine-gated monoatomic cation-selective channel activity ...sensory processing / synaptic transmission involved in micturition / response to acetylcholine / dendrite arborization / Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / acetylcholine receptor activity / acetylcholine-gated channel complex / chloride channel regulator activity / regulation of amyloid fibril formation / acetylcholine-gated monoatomic cation-selective channel activity / short-term memory / acetylcholine binding / dendritic spine organization / regulation of amyloid precursor protein catabolic process / acetylcholine receptor signaling pathway / positive regulation of amyloid-beta formation / negative regulation of amyloid-beta formation / response to amyloid-beta / positive regulation of protein metabolic process / negative regulation of tumor necrosis factor production / modulation of excitatory postsynaptic potential / plasma membrane raft / monoatomic ion channel activity / toxic substance binding / monoatomic ion transport / negative regulation of cytokine production involved in inflammatory response / positive regulation of excitatory postsynaptic potential / positive regulation of long-term synaptic potentiation / negative regulation of canonical NF-kappaB signal transduction / response to nicotine / regulation of membrane potential / excitatory postsynaptic potential / synapse organization / cognition / calcium channel activity / positive regulation of angiogenesis / memory / intracellular calcium ion homeostasis / calcium ion transport / transmembrane signaling receptor activity / amyloid-beta binding / monoatomic ion transmembrane transport / chemical synaptic transmission / learning or memory / response to hypoxia / positive regulation of MAPK cascade / positive regulation of ERK1 and ERK2 cascade / postsynaptic membrane / neuron projection / postsynapse / positive regulation of cell population proliferation / synapse / dendrite / endoplasmic reticulum membrane / signal transduction / protein homodimerization activity / membrane / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||||||||
 Authors Authors | Prevost MS / Barilone N / Dejean de la Batie G / Pons S / Ayme G / England P / Gielen M / Bontems F / Pehau-Arnaudet G / Maskos U ...Prevost MS / Barilone N / Dejean de la Batie G / Pons S / Ayme G / England P / Gielen M / Bontems F / Pehau-Arnaudet G / Maskos U / Lafaye P / Corringer P-J | ||||||||||||||||||
| Funding support | European Union,  France, 5 items France, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: An original potentiating mechanism revealed by the cryo-EM structures of the human α7 nicotinic receptor in complex with nanobodies. Authors: Marie S Prevost / Nathalie Barilone / Gabrielle Dejean de la Bâtie / Stéphanie Pons / Gabriel Ayme / Patrick England / Marc Gielen / François Bontems / Gérard Pehau-Arnaudet / Uwe Maskos ...Authors: Marie S Prevost / Nathalie Barilone / Gabrielle Dejean de la Bâtie / Stéphanie Pons / Gabriel Ayme / Patrick England / Marc Gielen / François Bontems / Gérard Pehau-Arnaudet / Uwe Maskos / Pierre Lafaye / Pierre-Jean Corringer /  Abstract: The human α7 nicotinic receptor is a pentameric channel mediating cellular and neuronal communication. It has attracted considerable interest in designing ligands for the treatment of neurological ...The human α7 nicotinic receptor is a pentameric channel mediating cellular and neuronal communication. It has attracted considerable interest in designing ligands for the treatment of neurological and psychiatric disorders. To develop a novel class of α7 ligands, we recently generated two nanobodies named E3 and C4, acting as positive allosteric modulator and silent allosteric ligand, respectively. Here, we solved the cryo-electron microscopy structures of the nanobody-receptor complexes. E3 and C4 bind to a common epitope involving two subunits at the apex of the receptor. They form by themselves a symmetric pentameric assembly that extends the extracellular domain. Unlike C4, the binding of E3 drives an agonist-bound conformation of the extracellular domain in the absence of an orthosteric agonist, and mutational analysis shows a key contribution of an N-linked sugar moiety in mediating E3 potentiation. The nanobody E3, by remotely controlling the global allosteric conformation of the receptor, implements an original mechanism of regulation that opens new avenues for drug design. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16534.map.gz emd_16534.map.gz | 80.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16534-v30.xml emd-16534-v30.xml emd-16534.xml emd-16534.xml | 20.5 KB 20.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16534.png emd_16534.png | 39.5 KB | ||
| Filedesc metadata |  emd-16534.cif.gz emd-16534.cif.gz | 6.8 KB | ||
| Others |  emd_16534_half_map_1.map.gz emd_16534_half_map_1.map.gz emd_16534_half_map_2.map.gz emd_16534_half_map_2.map.gz | 80.9 MB 80.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16534 http://ftp.pdbj.org/pub/emdb/structures/EMD-16534 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16534 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16534 | HTTPS FTP |
-Related structure data
| Related structure data |  8cauMC  8c9xC  8ce4C  8ci1C  8ci2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16534.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16534.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_16534_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
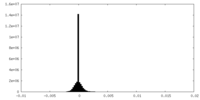
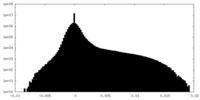
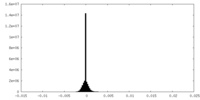
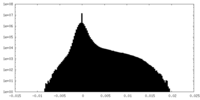
| Density Histograms |
-Half map: #1
| File | emd_16534_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of the human alpha7 nicotinic acetylcholine receptor with...
| Entire | Name: Complex of the human alpha7 nicotinic acetylcholine receptor withthe Nanobody C4 and with Nicotine |
|---|---|
| Components |
|
-Supramolecule #1: Complex of the human alpha7 nicotinic acetylcholine receptor with...
| Supramolecule | Name: Complex of the human alpha7 nicotinic acetylcholine receptor withthe Nanobody C4 and with Nicotine type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 250 KDa |
-Supramolecule #2: human alpha7 nicotinic acetylcholine receptor
| Supramolecule | Name: human alpha7 nicotinic acetylcholine receptor / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Nanobody C4
| Supramolecule | Name: Nanobody C4 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Neuronal acetylcholine receptor subunit alpha-7
| Macromolecule | Name: Neuronal acetylcholine receptor subunit alpha-7 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.36768 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EFQRKLYKEL VKNYNPLERP VANDSQPLTV YFSLSLLQIM DVDEKNQVLT TNIWLQMSWT DHYLQWNVSE YPGVKTVRFP DGQIWKPDI LLYNSADERF DATFHTNVLV NSSGHCQYLP PGIFKSSCYI DVRWFPFDVQ HCKLKFGSWS YGGWSLDLQM Q EADISGYI ...String: EFQRKLYKEL VKNYNPLERP VANDSQPLTV YFSLSLLQIM DVDEKNQVLT TNIWLQMSWT DHYLQWNVSE YPGVKTVRFP DGQIWKPDI LLYNSADERF DATFHTNVLV NSSGHCQYLP PGIFKSSCYI DVRWFPFDVQ HCKLKFGSWS YGGWSLDLQM Q EADISGYI PNGEWDLVGI PGKRSERFYE CCKEPYPDVT FTVTMRRRTL YYGLNLLIPC VLISALALLV FLLPADSGEK IS LGITVLL SLTVFMLLVA EIMPATSDSV PLIAQYFAST MIIVGLSVVV TVIVLQYHHH DPDGGKMPKW TRVILLNWCA WFL RMKRPG EDKVRPACQH KQRRCSLASV EMSAVAPPPA SNGNLLYIGF RGLDGVHCVP TPDSGVVCGR MACSPTHDEH LLHG GQPPE GDPDLAKILE EVRYIANRFR CQDESEAVCS EWKFAACVVD RLCLMAFSVF TIICTIGILM SAPNFVEAVS KDFAS AGLT ETSQVAPA UniProtKB: Neuronal acetylcholine receptor subunit alpha-7 |
-Macromolecule #2: Nanobody C4
| Macromolecule | Name: Nanobody C4 / type: protein_or_peptide / ID: 2 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.826398 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AQVQLVESGG GLVQAGGSLK LSCAASGFTF AHYAMVWFRQ APGKEREFVA GISWSGASTY YASSVKGRFT ISRDNAKNTV YLQMNSLKP EDTAVYYVAA ARFGVGVDDD YSYWGQGTQV TVSSAAEQKL ISEEDLNGAA HHHHHHGS |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 10 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: (S)-3-(1-METHYLPYRROLIDIN-2-YL)PYRIDINE
| Macromolecule | Name: (S)-3-(1-METHYLPYRROLIDIN-2-YL)PYRIDINE / type: ligand / ID: 5 / Number of copies: 5 / Formula: NCT |
|---|---|
| Molecular weight | Theoretical: 162.232 Da |
| Chemical component information |  ChemComp-NCT: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)