+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli BAM complex (BamABCDE) bound to darobactin B | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Outer Membrane Protein / Protein Folding / Antibiotic / beta barrel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationBam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface ...Bam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |   Photorhabdus heterorhabditis (bacteria) Photorhabdus heterorhabditis (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Horne JE / Fenn KL / Radford SE / Ranson NA | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Angew Chem Int Ed Engl / Year: 2023 Journal: Angew Chem Int Ed Engl / Year: 2023Title: Darobactin B Stabilises a Lateral-Closed Conformation of the BAM Complex in E. coli Cells. Authors: Samuel F Haysom / Jonathan Machin / James M Whitehouse / Jim E Horne / Katherine Fenn / Yue Ma / Hassane El Mkami / Nils Böhringer / Till F Schäberle / Neil A Ranson / Sheena E Radford / Christos Pliotas /   Abstract: The β-barrel assembly machinery (BAM complex) is essential for outer membrane protein (OMP) folding in Gram-negative bacteria, and represents a promising antimicrobial target. Several conformational ...The β-barrel assembly machinery (BAM complex) is essential for outer membrane protein (OMP) folding in Gram-negative bacteria, and represents a promising antimicrobial target. Several conformational states of BAM have been reported, but all have been obtained under conditions which lack the unique features and complexity of the outer membrane (OM). Here, we use Pulsed Electron-Electron Double Resonance (PELDOR, or DEER) spectroscopy distance measurements to interrogate the conformational ensemble of the BAM complex in E. coli cells. We show that BAM adopts a broad ensemble of conformations in the OM, while in the presence of the antibiotic darobactin B (DAR-B), BAM's conformational equilibrium shifts to a restricted ensemble consistent with the lateral closed state. Our in-cell PELDOR findings are supported by new cryoEM structures of BAM in the presence and absence of DAR-B. This work demonstrates the utility of PELDOR to map conformational changes in BAM within its native cellular environment. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16268.map.gz emd_16268.map.gz | 67 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16268-v30.xml emd-16268-v30.xml emd-16268.xml emd-16268.xml | 22.9 KB 22.9 KB | Display Display |  EMDB header EMDB header |
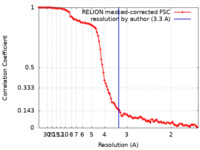
| FSC (resolution estimation) |  emd_16268_fsc.xml emd_16268_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_16268.png emd_16268.png | 152.3 KB | ||
| Others |  emd_16268_half_map_1.map.gz emd_16268_half_map_1.map.gz emd_16268_half_map_2.map.gz emd_16268_half_map_2.map.gz | 96.5 MB 96.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16268 http://ftp.pdbj.org/pub/emdb/structures/EMD-16268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16268 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16268 | HTTPS FTP |
-Related structure data
| Related structure data |  8bvqMC  8bwcC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16268.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16268.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_16268_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16268_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli BAM complex (BamABCDE) bound to darobactin B
| Entire | Name: E. coli BAM complex (BamABCDE) bound to darobactin B |
|---|---|
| Components |
|
-Supramolecule #1: E. coli BAM complex (BamABCDE) bound to darobactin B
| Supramolecule | Name: E. coli BAM complex (BamABCDE) bound to darobactin B / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 200.98298 KDa |
-Macromolecule #1: Outer membrane protein assembly factor BamA
| Macromolecule | Name: Outer membrane protein assembly factor BamA / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 88.514742 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVKE RPTIASITFS GNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP RNRVDLKLVF QEGVSAEIQQ I NIVGNHAF ...String: AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVKE RPTIASITFS GNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP RNRVDLKLVF QEGVSAEIQQ I NIVGNHAF TTDELISHFQ LRDEVPWWNV VGDRKYQKQK LAGDLETLRS YYLDRGYARF NIDSTQVSLT PDKKGIYVTV NI TEGDQYK LSGVEVSGNL AGHSAEIEQL TKIEPGELYN GTKVTKMEDD IKKLLGRYGY AYPRVQSMPE INDADKTVKL RVN VDAGNR FYVRKIRFEG NDTSKDAVLR REMRQMEGAW LGSDLVDQGK ERLNRLGFFE TVDTDTQRVP GSPDQVDVVY KVKE RNTGS FNFGIGYGTE SGVSFQAGVQ QDNWLGTGYA VGINGTKNDY QTYAELSVTN PYFTVDGVSL GGRLFYNDFQ ADDAD LSDY TNKSYGTDVT LGFPINEYNS LRAGLGYVHN SLSNMQPQVA MWRYLYSMGE HPSTSDQDNS FKTDDFTFNY GWTYNK LDR GYFPTDGSRV NLTGKVTIPG SDNEYYKVTL DTATYVPIDD DHKWVVLGRT RWGYGDGLGG KEMPFYENFY AGGSSTV RG FQSNTIGPKA VYFPHQASNY DPDYDYECAT QDGAKDLCKS DDAVGGNAMA VASLEFITPT PFISDKYANS VRTSFFWD M GTVWDTNWDS SQYSGYPDYS DPSNIRMSAG IALQWMSPLG PLVFSYAQPF KKYDGDKAEQ FQFNIGKTW UniProtKB: Outer membrane protein assembly factor BamA |
-Macromolecule #2: Outer membrane protein assembly factor BamB
| Macromolecule | Name: Outer membrane protein assembly factor BamB / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.882375 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CSLFNSEEDV VKMSPLPTVE NQFTPTTAWS TSVGSGIGNF YSNLHPALAD NVVYAADRAG LVKALNADDG KEIWSVSLAE KDGWFSKEP ALLSGGVTVS GGHVYIGSEK AQVYALNTSD GTVAWQTKVA GEALSRPVVS DGLVLIHTSN GQLQALNEAD G AVKWTVNL ...String: CSLFNSEEDV VKMSPLPTVE NQFTPTTAWS TSVGSGIGNF YSNLHPALAD NVVYAADRAG LVKALNADDG KEIWSVSLAE KDGWFSKEP ALLSGGVTVS GGHVYIGSEK AQVYALNTSD GTVAWQTKVA GEALSRPVVS DGLVLIHTSN GQLQALNEAD G AVKWTVNL DMPSLSLRGE SAPTTAFGAA VVGGDNGRVS AVLMEQGQMI WQQRISQATG STEIDRLSDV DTTPVVVNGV VF ALAYNGN LTALDLRSGQ IMWKRELGSV NDFIVDGNRI YLVDQNDRVM ALTIDGGVTL WTQSDLLHRL LTSPVLYNGN LVV GDSEGY LHWINVEDGR FVAQQKVDSS GFQTEPVAAD GKLLIQAKDG TVYSITR UniProtKB: Outer membrane protein assembly factor BamB |
-Macromolecule #3: Outer membrane protein assembly factor BamC
| Macromolecule | Name: Outer membrane protein assembly factor BamC / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.40125 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CSSDSRYKRQ VSGDEAYLEA APLAELHAPA GMILPVTSGD YAIPVTNGSG AVGKALDIRP PAQPLALVSG ARTQFTGDTA SLLVENGRG NTLWPQVVSV LQAKNYTITQ RDDAGQTLTT DWVQWNRLDE DEQYRGRYQI SVKPQGYQQA VTVKLLNLEQ A GKPVADAA ...String: CSSDSRYKRQ VSGDEAYLEA APLAELHAPA GMILPVTSGD YAIPVTNGSG AVGKALDIRP PAQPLALVSG ARTQFTGDTA SLLVENGRG NTLWPQVVSV LQAKNYTITQ RDDAGQTLTT DWVQWNRLDE DEQYRGRYQI SVKPQGYQQA VTVKLLNLEQ A GKPVADAA SMQRYSTEMM NVISAGLDKS ATDAANAAQN RASTTMDVQS AADDTGLPML VVRGPFNVVW QRLPAALEKV GM KVTDSTR SQGNMAVTYK PLSDSDWQEL GASDPGLASG DYKLQVGDLD NRSSLQFIDP KGHTLTQSQN DALVAVFQAA FSK UniProtKB: Outer membrane protein assembly factor BamC |
-Macromolecule #4: Outer membrane protein assembly factor BamD
| Macromolecule | Name: Outer membrane protein assembly factor BamD / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.816818 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CSGSKEEVPD NPPNEIYATA QQKLQDGNWR QAITQLEALD NRYPFGPYSQ QVQLDLIYAY YKNADLPLAQ AAIDRFIRLN PTHPNIDYV MYMRGLTNMA LDDSALQGFF GVDRSDRDPQ HARAAFSDFS KLVRGYPNSQ YTTDATKRLV FLKDRLAKYE Y SVAEYYTE ...String: CSGSKEEVPD NPPNEIYATA QQKLQDGNWR QAITQLEALD NRYPFGPYSQ QVQLDLIYAY YKNADLPLAQ AAIDRFIRLN PTHPNIDYV MYMRGLTNMA LDDSALQGFF GVDRSDRDPQ HARAAFSDFS KLVRGYPNSQ YTTDATKRLV FLKDRLAKYE Y SVAEYYTE RGAWVAVVNR VEGMLRDYPD TQATRDALPL MENAYRQMQM NAQAEKVAKI IAANSSNT UniProtKB: Outer membrane protein assembly factor BamD |
-Macromolecule #5: Outer membrane protein assembly factor BamE
| Macromolecule | Name: Outer membrane protein assembly factor BamE / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.610833 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CSTLERVVYR PDINQGNYLT ANDVSKIRVG MTQQQVAYAL GTPLMSDPFG TNTWFYVFRQ QPGHEGVTQQ TLTLTFNSSG VLTNIDNKP ALSGNGGHHH HHHHH UniProtKB: Outer membrane protein assembly factor BamE |
-Macromolecule #6: Darobactin-B
| Macromolecule | Name: Darobactin-B / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Photorhabdus heterorhabditis (bacteria) Photorhabdus heterorhabditis (bacteria) |
| Molecular weight | Theoretical: 1.055189 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: WN(UX8)TKRF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 3732 / Average electron dose: 34.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8bvq: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)