[English] 日本語
 Yorodumi
Yorodumi- EMDB-16221: Triple-stranded DNA as a structural element in DNA origami rod object -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Triple-stranded DNA as a structural element in DNA origami rod object | |||||||||||||||
 Map data Map data | 10 helix bundle DNA origami object | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | DNA Origami / Triplex forming Oligo / Triplex / TFO / DNA | |||||||||||||||
| Biological species | synthetic construct (others) /  Phage M13mp18 (virus) Phage M13mp18 (virus) | |||||||||||||||
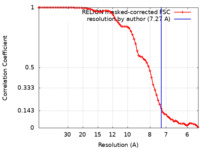
| Method | single particle reconstruction / cryo EM / Resolution: 7.27 Å | |||||||||||||||
 Authors Authors | Sachenbacher K / Khoshouei A / Honemann MN / Engelen W / Feigl E / Dietz H | |||||||||||||||
| Funding support | European Union,  Germany, Germany,  Netherlands, 4 items Netherlands, 4 items
| |||||||||||||||
 Citation Citation |  Journal: ACS Nano / Year: 2023 Journal: ACS Nano / Year: 2023Title: Triple-Stranded DNA As a Structural Element in DNA Origami. Authors: Ken Sachenbacher / Ali Khoshouei / Maximilian Nicolas Honemann / Wouter Engelen / Elija Feigl / Hendrik Dietz /  Abstract: Molecular self-assembly with DNA origami offers an attractive route to fabricate arbitrary three-dimensional nanostructures. In DNA origami, B-form double-helical DNA domains (dsDNA) are commonly ...Molecular self-assembly with DNA origami offers an attractive route to fabricate arbitrary three-dimensional nanostructures. In DNA origami, B-form double-helical DNA domains (dsDNA) are commonly linked with covalent phosphodiester strand crossovers to build up three-dimensional objects. To expand the palette of structural motifs in DNA origami, here we describe hybrid duplex-triplex DNA motifs as pH-dependent building blocks in DNA origami. We investigate design rules for incorporating triplex forming oligonucleotides and noncanonical duplex-triplex crossovers in multilayer DNA origami objects. We use single-particle cryoelectron microscopy to elucidate the structural basis of triplex domains and of duplex-triplex crossovers. We find that duplex-triplex crossovers can complement and fully replace the canonical duplex-duplex crossovers within DNA origami objects, for example, to increase the crossover density for potentially greater rigidity and reduced interhelical spacing, and to create connections at sites where conventional crossovers may be undesirable. We also show the pH-induced formation of a DNA origami object stabilized entirely by triplex-mediated strand crossovers. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16221.map.gz emd_16221.map.gz | 3.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16221-v30.xml emd-16221-v30.xml emd-16221.xml emd-16221.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16221_fsc.xml emd_16221_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_16221.png emd_16221.png | 67.1 KB | ||
| Others |  emd_16221_half_map_1.map.gz emd_16221_half_map_1.map.gz emd_16221_half_map_2.map.gz emd_16221_half_map_2.map.gz | 23.4 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16221 http://ftp.pdbj.org/pub/emdb/structures/EMD-16221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16221 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16221 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16221.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16221.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 10 helix bundle DNA origami object | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.8 Å | ||||||||||||||||||||||||||||||||||||
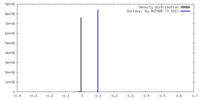
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: 10 helix bundle DNA origami object / half map 1
| File | emd_16221_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 10 helix bundle DNA origami object / half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
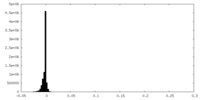
| Density Histograms |
-Half map: 10 helix bundle DNA origami object / half map 2
| File | emd_16221_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 10 helix bundle DNA origami object / half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
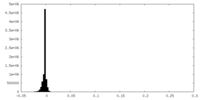
| Density Histograms |
- Sample components
Sample components
-Entire : DNA Nanostructure
| Entire | Name: DNA Nanostructure |
|---|---|
| Components |
|
-Supramolecule #1: DNA Nanostructure
| Supramolecule | Name: DNA Nanostructure / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Supramolecule #2: Scaffold DNA single-strand
| Supramolecule | Name: Scaffold DNA single-strand / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Phage M13mp18 (virus) Phage M13mp18 (virus) |
-Supramolecule #3: staple oligonucleotides
| Supramolecule | Name: staple oligonucleotides / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 5 / Details: 5 mM MES, 5mM NaCl and 5 mM MgCl2 |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 47000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)