+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | FcMR binding at subunit Fcu1 of IgM pentamer | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | IgM Fc receptor / IMMUNE SYSTEM | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationhigh-affinity IgM receptor activity / immunoglobulin transcytosis in epithelial cells / IgM binding / regulation of B cell receptor signaling pathway / polymeric immunoglobulin binding / Fc receptor-mediated immune complex endocytosis / humoral immune response mediated by circulating immunoglobulin / cellular defense response / trans-Golgi network membrane / transmembrane signaling receptor activity ...high-affinity IgM receptor activity / immunoglobulin transcytosis in epithelial cells / IgM binding / regulation of B cell receptor signaling pathway / polymeric immunoglobulin binding / Fc receptor-mediated immune complex endocytosis / humoral immune response mediated by circulating immunoglobulin / cellular defense response / trans-Golgi network membrane / transmembrane signaling receptor activity / early endosome membrane / lysosomal membrane / centrosome / negative regulation of apoptotic process / Golgi apparatus / signal transduction / extracellular region / nucleoplasm / plasma membrane Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||||||||||||||
 Authors Authors | Chen Q / Rosenthal P / Tolar P | |||||||||||||||||||||
| Funding support |  United Kingdom, 6 items United Kingdom, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structural basis for Fc receptor recognition of immunoglobulin M. Authors: Qu Chen / Rajesh P Menon / Laura Masino / Pavel Tolar / Peter B Rosenthal /  Abstract: Immunoglobulin Fc receptors are cell surface transmembrane proteins that bind to the Fc constant region of antibodies and play critical roles in regulating immune responses by activation of immune ...Immunoglobulin Fc receptors are cell surface transmembrane proteins that bind to the Fc constant region of antibodies and play critical roles in regulating immune responses by activation of immune cells, clearance of immune complexes and regulation of antibody production. FcμR is the immunoglobulin M (IgM) antibody isotype-specific Fc receptor involved in the survival and activation of B cells. Here we reveal eight binding sites for the human FcμR immunoglobulin domain on the IgM pentamer by cryogenic electron microscopy. One of the sites overlaps with the binding site for the polymeric immunoglobulin receptor (pIgR), but a different mode of FcμR binding explains its antibody isotype specificity. Variation in FcμR binding sites and their occupancy reflects the asymmetry of the IgM pentameric core and the versatility of FcμR binding. The complex explains engagement with polymeric serum IgM and the monomeric IgM B-cell receptor (BCR). | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16151.map.gz emd_16151.map.gz | 16.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16151-v30.xml emd-16151-v30.xml emd-16151.xml emd-16151.xml | 22.4 KB 22.4 KB | Display Display |  EMDB header EMDB header |
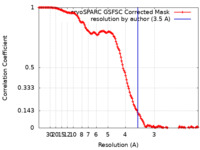
| FSC (resolution estimation) |  emd_16151_fsc.xml emd_16151_fsc.xml | 15 KB | Display |  FSC data file FSC data file |
| Images |  emd_16151.png emd_16151.png | 88.2 KB | ||
| Masks |  emd_16151_msk_1.map emd_16151_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16151.cif.gz emd-16151.cif.gz | 7.1 KB | ||
| Others |  emd_16151_half_map_1.map.gz emd_16151_half_map_1.map.gz emd_16151_half_map_2.map.gz emd_16151_half_map_2.map.gz | 226.9 MB 226.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16151 http://ftp.pdbj.org/pub/emdb/structures/EMD-16151 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16151 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16151 | HTTPS FTP |
-Validation report
| Summary document |  emd_16151_validation.pdf.gz emd_16151_validation.pdf.gz | 679.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16151_full_validation.pdf.gz emd_16151_full_validation.pdf.gz | 678.8 KB | Display | |
| Data in XML |  emd_16151_validation.xml.gz emd_16151_validation.xml.gz | 22.1 KB | Display | |
| Data in CIF |  emd_16151_validation.cif.gz emd_16151_validation.cif.gz | 28.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16151 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16151 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16151 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16151 | HTTPS FTP |
-Related structure data
| Related structure data |  8bpfMC  8bpeC  8bpgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16151.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16151.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16151_msk_1.map emd_16151_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16151_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16151_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human FcMR/IgM-Fc complex
| Entire | Name: human FcMR/IgM-Fc complex |
|---|---|
| Components |
|
-Supramolecule #1: human FcMR/IgM-Fc complex
| Supramolecule | Name: human FcMR/IgM-Fc complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1, #3, #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Immunoglobulin heavy constant mu
| Macromolecule | Name: Immunoglobulin heavy constant mu / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.343086 KDa |
| Sequence | String: IAELPPKVSV FVPPRDGFFG NPRKSKLICQ ATGFSPRQIQ VSWLREGKQV GSGVTTDQVQ AEAKESGPTT YKVTSTLTIK ESDWLGQSM FTCRVDHRGL TFQQNASSMC VPDQDTAIRV FAIPPSFASI FLTKSTKLTC LVTDLTTYDS VTISWTRQNG E AVKTHTNI ...String: IAELPPKVSV FVPPRDGFFG NPRKSKLICQ ATGFSPRQIQ VSWLREGKQV GSGVTTDQVQ AEAKESGPTT YKVTSTLTIK ESDWLGQSM FTCRVDHRGL TFQQNASSMC VPDQDTAIRV FAIPPSFASI FLTKSTKLTC LVTDLTTYDS VTISWTRQNG E AVKTHTNI SESHPNATFS AVGEASICED DWNSGERFTC TVTHTDLPSP LKQTISRPKG VALHRPDVYL LPPAREQLNL RE SATITCL VTGFSPADVF VQWMQRGQPL SPEKYVTSAP MPEPQAPGRY FAHSILTVSE EEWNTGETYT CVVAHEALPN RVT ERTVDK STGKPTLYNV SLVMSDTAGT CY |
-Macromolecule #2: Fas apoptotic inhibitory molecule 3
| Macromolecule | Name: Fas apoptotic inhibitory molecule 3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.020617 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: RILPEVKVEG ELGGSVTIKC PLPEMHVRIY LCREMAGSGT CGTVVSTTNF IKAEYKGRVT LKQYPRKNLF LVEVTQLTES DSGVYACGA GMNTDRGKTQ KVTLNVHSEY EPSWEEQPMP ETPKWFHLPY LFQMPAYASS SKFVTRVTTP AQRGKVPPVH H SSPTTQIT ...String: RILPEVKVEG ELGGSVTIKC PLPEMHVRIY LCREMAGSGT CGTVVSTTNF IKAEYKGRVT LKQYPRKNLF LVEVTQLTES DSGVYACGA GMNTDRGKTQ KVTLNVHSEY EPSWEEQPMP ETPKWFHLPY LFQMPAYASS SKFVTRVTTP AQRGKVPPVH H SSPTTQIT HRPRVSRASS VAGDKPRTFL PSTTASKISA LEGLLKPQTP SYNHHTRLHR QRALDYGSQS GREGQG UniProtKB: Immunoglobulin mu Fc receptor |
-Macromolecule #3: Immunoglobulin J chain
| Macromolecule | Name: Immunoglobulin J chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.120586 KDa |
| Sequence | String: MKNHLLFWGV LAVFIKAVHV KAQEDERIVL VDNKCKCARI TSRIIRSSED PNEDIVERNI RIIVPLNNRE NISDPTSPLR TRFVYHLSD LCKKCDPTEV ELDNQIVTAT QSNICDEDSA TETCYTYDRN KCYTAVVPLV YGGETKMVET ALTPDACYPD |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 10 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 4.0 µm / Calibrated defocus min: 1.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8bpf: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)