[English] 日本語
 Yorodumi
Yorodumi- EMDB-15966: Structure of Synechococcus elongatus PCC 7942 Rubisco recombinant... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Synechococcus elongatus PCC 7942 Rubisco recombinantly expressed from E.coli | |||||||||
 Map data Map data | main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Rubisco / Synechococcus / carbon dioxide fixation / recombinant expression / BIOSYNTHETIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationphotorespiration / carboxysome / ribulose-bisphosphate carboxylase / ribulose-bisphosphate carboxylase activity / reductive pentose-phosphate cycle / monooxygenase activity / magnesium ion binding Similarity search - Function | |||||||||
| Biological species |  Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.15 Å | |||||||||
 Authors Authors | Ni T / Sun Y / Liu LN / Zhang P | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of Rubisco Authors: Sun Y / Ni T | |||||||||
| History |
|
- Structure visualization
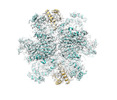
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15966.map.gz emd_15966.map.gz | 93.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15966-v30.xml emd-15966-v30.xml emd-15966.xml emd-15966.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15966.png emd_15966.png | 117 KB | ||
| Filedesc metadata |  emd-15966.cif.gz emd-15966.cif.gz | 5.9 KB | ||
| Others |  emd_15966_half_map_1.map.gz emd_15966_half_map_1.map.gz emd_15966_half_map_2.map.gz emd_15966_half_map_2.map.gz | 91.8 MB 91.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15966 http://ftp.pdbj.org/pub/emdb/structures/EMD-15966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15966 | HTTPS FTP |
-Related structure data
| Related structure data |  8bcmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15966.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15966.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 1
| File | emd_15966_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_15966_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Synechococcus elongatus PCC 7942 Rubisco recombinantly co-express...
| Entire | Name: Synechococcus elongatus PCC 7942 Rubisco recombinantly co-expressed in E.coli |
|---|---|
| Components |
|
-Supramolecule #1: Synechococcus elongatus PCC 7942 Rubisco recombinantly co-express...
| Supramolecule | Name: Synechococcus elongatus PCC 7942 Rubisco recombinantly co-expressed in E.coli type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) |
-Macromolecule #1: Ribulose 1,5-bisphosphate carboxylase small subunit
| Macromolecule | Name: Ribulose 1,5-bisphosphate carboxylase small subunit / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:  Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria)Strain: PCC 7942 |
| Molecular weight | Theoretical: 13.349196 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSMKTLPKER RFETFSYLPP LSDRQIAAQI EYMIEQGFHP LIEFNEHSNP EEFYWTMWKL PLFDCKSPQQ VLDEVRECRS EYGDCYIRV AGFDNIKQCQ TVSFIVHRPG RY UniProtKB: Ribulose bisphosphate carboxylase small subunit |
-Macromolecule #2: Ribulose bisphosphate carboxylase large chain
| Macromolecule | Name: Ribulose bisphosphate carboxylase large chain / type: protein_or_peptide / ID: 2 / Number of copies: 8 / Enantiomer: LEVO / EC number: ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:  Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria) Synechococcus elongatus PCC 7942 = FACHB-805 (bacteria)Strain: PCC 7942 |
| Molecular weight | Theoretical: 52.516605 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPKTQSAAGY KAGVKDYKLT YYTPDYTPKD TDLLAAFRFS PQPGVPADEA GAAIAAESST GTWTTVWTDL LTDMDRYKGK CYHIEPVQG EENSYFAFIA YPLDLFEEGS VTNILTSIVG NVFGFKAIRS LRLEDIRFPV ALVKTFQGPP HGIQVERDLL N KYGRPMLG ...String: MPKTQSAAGY KAGVKDYKLT YYTPDYTPKD TDLLAAFRFS PQPGVPADEA GAAIAAESST GTWTTVWTDL LTDMDRYKGK CYHIEPVQG EENSYFAFIA YPLDLFEEGS VTNILTSIVG NVFGFKAIRS LRLEDIRFPV ALVKTFQGPP HGIQVERDLL N KYGRPMLG CTIKPKLGLS AKNYGRAVYE CLRGGLDFTK DDENINSQPF QRWRDRFLFV ADAIHKSQAE TGEIKGHYLN VT APTCEEM MKRAEFAKEL GMPIIMHDFL TAGFTANTTL AKWCRDNGVL LHIHRAMHAV IDRQRNHGIH FRVLAKCLRL SGG DHLHSG TVVGKLEGDK ASTLGFVDLM REDHIEADRS RGVFFTQDWA SMPGVLPVAS GGIHVWHMPA LVEIFGDDSV LQFG GGTLG HPWGNAPGAT ANRVALEACV QARNEGRDLY REGGDILREA GKWSPELAAA LDLWKEIKFE FETMDKL UniProtKB: Ribulose bisphosphate carboxylase large chain |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8bcm: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)