[English] 日本語
 Yorodumi
Yorodumi- EMDB-15775: Cryo-EM structure of the Tripartite ATP-independent Periplasmic (... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Photobacterium profundum in a nanodisc | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRAP / elevator / secondary transporter / sialic acid / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationC4-dicarboxylate transport / transmembrane transporter activity / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Photobacterium profundum (bacteria) / Photobacterium profundum (bacteria) /   Photobacterium profundum SS9 (bacteria) / synthetic construct (others) Photobacterium profundum SS9 (bacteria) / synthetic construct (others) | |||||||||
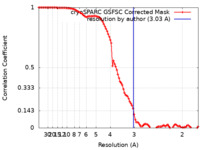
| Method | single particle reconstruction / cryo EM / Resolution: 3.03 Å | |||||||||
 Authors Authors | Davies JS / North RA / Dobson RCJ | |||||||||
| Funding support |  New Zealand, 2 items New Zealand, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structure and mechanism of a tripartite ATP-independent periplasmic TRAP transporter. Authors: James S Davies / Michael J Currie / Rachel A North / Mariafrancesca Scalise / Joshua D Wright / Jack M Copping / Daniela M Remus / Ashutosh Gulati / Dustin R Morado / Sam A Jamieson / ...Authors: James S Davies / Michael J Currie / Rachel A North / Mariafrancesca Scalise / Joshua D Wright / Jack M Copping / Daniela M Remus / Ashutosh Gulati / Dustin R Morado / Sam A Jamieson / Michael C Newton-Vesty / Gayan S Abeysekera / Subramanian Ramaswamy / Rosmarie Friemann / Soichi Wakatsuki / Jane R Allison / Cesare Indiveri / David Drew / Peter D Mace / Renwick C J Dobson /      Abstract: In bacteria and archaea, tripartite ATP-independent periplasmic (TRAP) transporters uptake essential nutrients. TRAP transporters receive their substrates via a secreted soluble substrate-binding ...In bacteria and archaea, tripartite ATP-independent periplasmic (TRAP) transporters uptake essential nutrients. TRAP transporters receive their substrates via a secreted soluble substrate-binding protein. How a sodium ion-driven secondary active transporter is strictly coupled to a substrate-binding protein is poorly understood. Here we report the cryo-EM structure of the sialic acid TRAP transporter SiaQM from Photobacterium profundum at 2.97 Å resolution. SiaM comprises a "transport" domain and a "scaffold" domain, with the transport domain consisting of helical hairpins as seen in the sodium ion-coupled elevator transporter VcINDY. The SiaQ protein forms intimate contacts with SiaM to extend the size of the scaffold domain, suggesting that TRAP transporters may operate as monomers, rather than the typically observed oligomers for elevator-type transporters. We identify the Na and sialic acid binding sites in SiaM and demonstrate a strict dependence on the substrate-binding protein SiaP for uptake. We report the SiaP crystal structure that, together with docking studies, suggest the molecular basis for how sialic acid is delivered to the SiaQM transporter complex. We thus propose a model for substrate transport by TRAP proteins, which we describe herein as an 'elevator-with-an-operator' mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15775.map.gz emd_15775.map.gz | 51.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15775-v30.xml emd-15775-v30.xml emd-15775.xml emd-15775.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15775_fsc.xml emd_15775_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_15775.png emd_15775.png | 92.9 KB | ||
| Filedesc metadata |  emd-15775.cif.gz emd-15775.cif.gz | 6.7 KB | ||
| Others |  emd_15775_half_map_1.map.gz emd_15775_half_map_1.map.gz emd_15775_half_map_2.map.gz emd_15775_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15775 http://ftp.pdbj.org/pub/emdb/structures/EMD-15775 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15775 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15775 | HTTPS FTP |
-Validation report
| Summary document |  emd_15775_validation.pdf.gz emd_15775_validation.pdf.gz | 919.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15775_full_validation.pdf.gz emd_15775_full_validation.pdf.gz | 918.7 KB | Display | |
| Data in XML |  emd_15775_validation.xml.gz emd_15775_validation.xml.gz | 18.2 KB | Display | |
| Data in CIF |  emd_15775_validation.cif.gz emd_15775_validation.cif.gz | 23.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15775 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15775 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15775 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15775 | HTTPS FTP |
-Related structure data
| Related structure data |  8b01MC  7qhaC  7t3eC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15775.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15775.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.886 Å | ||||||||||||||||||||||||||||||||||||
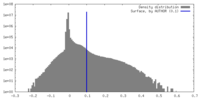
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_15775_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
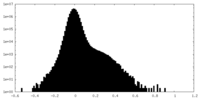
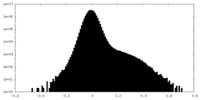
| Density Histograms |
-Half map: #1
| File | emd_15775_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
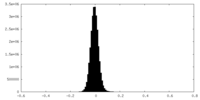
| Density Histograms |
- Sample components
Sample components
+Entire : SiaQM with megabody
+Supramolecule #1: SiaQM with megabody
+Supramolecule #2: SiaQM
+Supramolecule #3: MegaBody
+Macromolecule #1: Putative TRAP-type C4-dicarboxylate transport system, small perme...
+Macromolecule #2: Putative TRAP-type C4-dicarboxylate transport system, large perme...
+Macromolecule #3: Megabody c7HopQ
+Macromolecule #4: N-OCTANE
+Macromolecule #5: PHOSPHATIDYLETHANOLAMINE
+Macromolecule #6: DOCOSANE
+Macromolecule #7: DECANE
+Macromolecule #8: SODIUM ION
+Macromolecule #9: TRIDECANE
+Macromolecule #10: HEXANE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 8127 / Average exposure time: 2.0 sec. / Average electron dose: 70.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.4 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)