+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | myo-Inositol-1-Phosphate Synthase | |||||||||||||||||||||||||||
 Map data Map data | Full map of the reconstructed myo-inositol-1-phosphate synthase | |||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||
| Biological species |  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) | |||||||||||||||||||||||||||
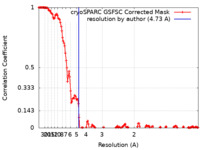
| Method | single particle reconstruction / cryo EM / Resolution: 4.73 Å | |||||||||||||||||||||||||||
 Authors Authors | Janson K / Kyrilis FL / Tueting C / Alfes M / Das M / Traeger TK / Schmidt C / Hamdi F / Keller S / Meister A / Kastritis PL | |||||||||||||||||||||||||||
| Funding support |  Germany, European Union, Germany, European Union,  France, France,  Austria, 8 items Austria, 8 items
| |||||||||||||||||||||||||||
 Citation Citation |  Journal: Biomacromolecules / Year: 2022 Journal: Biomacromolecules / Year: 2022Title: Cryo-Electron Microscopy Snapshots of Eukaryotic Membrane Proteins in Native Lipid-Bilayer Nanodiscs. Authors: Kevin Janson / Fotis L Kyrilis / Christian Tüting / Marie Alfes / Manabendra Das / Toni K Träger / Carla Schmidt / Farzad Hamdi / Carolyn Vargas / Sandro Keller / Annette Meister / Panagiotis L Kastritis /   Abstract: New technologies for purifying membrane-bound protein complexes in combination with cryo-electron microscopy (EM) have recently allowed the exploration of such complexes under near-native conditions. ...New technologies for purifying membrane-bound protein complexes in combination with cryo-electron microscopy (EM) have recently allowed the exploration of such complexes under near-native conditions. In particular, polymer-encapsulated nanodiscs enable the study of membrane proteins at high resolution while retaining protein-protein and protein-lipid interactions within a lipid bilayer. However, this powerful technology has not been exploited to address the important question of how endogenous─as opposed to overexpressed─membrane proteins are organized within a lipid environment. In this work, we demonstrate that biochemical enrichment protocols for native membrane-protein complexes from in combination with polymer-based lipid-bilayer nanodiscs provide a substantial improvement in the quality of recovered endogenous membrane-protein complexes. Mass spectrometry results revealed ∼1123 proteins, while multiple 2D class averages and two 3D reconstructions from cryo-EM data furnished prominent structural signatures. This integrated methodological approach to enriching endogenous membrane-protein complexes provides unprecedented opportunities for a deeper understanding of eukaryotic membrane proteomes. | |||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15517.map.gz emd_15517.map.gz | 323.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15517-v30.xml emd-15517-v30.xml emd-15517.xml emd-15517.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15517_fsc.xml emd_15517_fsc.xml | 15 KB | Display |  FSC data file FSC data file |
| Images |  emd_15517.png emd_15517.png | 54.1 KB | ||
| Masks |  emd_15517_msk_1.map emd_15517_msk_1.map | 343 MB |  Mask map Mask map | |
| Others |  emd_15517_half_map_1.map.gz emd_15517_half_map_1.map.gz emd_15517_half_map_2.map.gz emd_15517_half_map_2.map.gz | 318.7 MB 318.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15517 http://ftp.pdbj.org/pub/emdb/structures/EMD-15517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15517 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15517 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15517.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15517.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map of the reconstructed myo-inositol-1-phosphate synthase | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.5918 Å | ||||||||||||||||||||||||||||||||||||
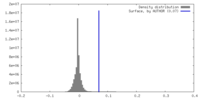
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15517_msk_1.map emd_15517_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
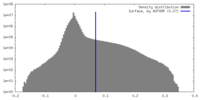
| Density Histograms |
-Half map: Half map of the reconstructed
| File | emd_15517_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of the reconstructed | ||||||||||||
| Projections & Slices |
| ||||||||||||
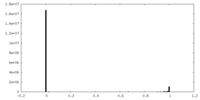
| Density Histograms |
-Half map: Half map of the reconstructed
| File | emd_15517_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of the reconstructed | ||||||||||||
| Projections & Slices |
| ||||||||||||
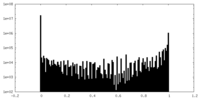
| Density Histograms |
- Sample components
Sample components
-Entire : SB-DIBMA solubilized Chaetomium thermophilum membranes mMW fraction
| Entire | Name: SB-DIBMA solubilized Chaetomium thermophilum membranes mMW fraction |
|---|---|
| Components |
|
-Supramolecule #1: SB-DIBMA solubilized Chaetomium thermophilum membranes mMW fraction
| Supramolecule | Name: SB-DIBMA solubilized Chaetomium thermophilum membranes mMW fraction type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all Details: Sample was created by solubilizing native chaetomium termophilum membranes with the aid of the copolymer SB-DIBMA |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) |
| Molecular weight | Theoretical: 120 KDa |
-Macromolecule #1: myo-Inositol-1-Phosphate Synthase
| Macromolecule | Name: myo-Inositol-1-Phosphate Synthase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thermochaetoides thermophila (fungus) Thermochaetoides thermophila (fungus) |
| Sequence | String: MAPHAEVDAG LANGGGQANG NGVAAAVAAP TVAPTTVSPI FKVNSPNVVY TDDEIRSKYV YRTTEVTTAE DGSLIATPRE TVYDFKVDRK LPKLGVMLVG WGGNNGSTIT AGIIANRRGL VWETRNGKQE ANYYGSVIMG STIKLGTDAK THKDINIPFH SVLPMVHPND ...String: MAPHAEVDAG LANGGGQANG NGVAAAVAAP TVAPTTVSPI FKVNSPNVVY TDDEIRSKYV YRTTEVTTAE DGSLIATPRE TVYDFKVDRK LPKLGVMLVG WGGNNGSTIT AGIIANRRGL VWETRNGKQE ANYYGSVIMG STIKLGTDAK THKDINIPFH SVLPMVHPND IVIGGWDISG LNLADAMDRA QVLEPSLKAL VRKEMASMKP LPSIYYPDFI AANQEDRADN ILPGNKACWE HVEEIRKNIR DFKAANGLDK VIVLWTANTE RYASIIEGVN DTADNLLNAI KNGHEEVSPS TVFAVSSILE GVPFINGSPQ NTFVPGCIEL AERHGAFIGG DDFKSGQTKM KSALVDFLIN AGIKLTSIAS YNHLGNNDGK NLSSQRQFRS KEISKSNVVD DMVEANTVLY KPGEHPDHIV VIKYVPAVGD SKRAMDEYHG EIFLGGHQTI SIANVCEDSL LASPLIIDLV IVAELMTRIQ WRLHKEDATE ADWKYFHSVL SILSYMLKAP MTPPGTPVVN ALAKQRAAMA NIFRACLGLD PENDMTLEHK LF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Solutions were freshly prepared, sterile filtrated, and sonicated before usage | |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.04 kPa Details: No special treatment The Grid was charged with 15 mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV / Details: Blotting time of 12 s Blotforce of 0. | |||||||||
| Details | This sample was purified by size exclusion chromatography. Subsequently, multiple fractions in the medium molecular weight region were pooled together to obtain a higher protein concentration. |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Temperature | Min: 77.15 K / Max: 103.15 K |
| Alignment procedure | Coma free - Residual tilt: 14.7 mrad |
| Details | Grid screening was performed manually until criteria for good acquisition areas was narrowed down. |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4048 pixel / Digitization - Dimensions - Height: 4048 pixel / Number grids imaged: 1 / Number real images: 5912 / Average electron dose: 64.72 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 253464 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 240000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)