[English] 日本語
 Yorodumi
Yorodumi- EMDB-15480: Subtomogram average of nucleosomes extracted from euchromatin and... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of nucleosomes extracted from euchromatin and facultative heterochromatin nanodomains of Drosophila melanogaster embryos | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | nucleosome tomography chromatin / STRUCTURAL GENOMICS | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 13.5 Å | |||||||||||||||
 Authors Authors | Fatmaoui F / Eltsov M / Leforestier A | |||||||||||||||
| Funding support |  France, France,  Germany, 4 items Germany, 4 items
| |||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-electron tomography and deep learning-based denoising reveal native chromatin landscapes of interphase nuclei Authors: Fatmaoui F / Carrivain P / Grewe D / Jakob B / Victor JM / Leforestier A / Eltsov M | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15480.map.gz emd_15480.map.gz | 1005.4 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15480-v30.xml emd-15480-v30.xml emd-15480.xml emd-15480.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15480.png emd_15480.png | 39 KB | ||
| Others |  emd_15480_half_map_1.map.gz emd_15480_half_map_1.map.gz emd_15480_half_map_2.map.gz emd_15480_half_map_2.map.gz | 1005.2 KB 1005.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15480 http://ftp.pdbj.org/pub/emdb/structures/EMD-15480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15480 | HTTPS FTP |
-Validation report
| Summary document |  emd_15480_validation.pdf.gz emd_15480_validation.pdf.gz | 508.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15480_full_validation.pdf.gz emd_15480_full_validation.pdf.gz | 508.3 KB | Display | |
| Data in XML |  emd_15480_validation.xml.gz emd_15480_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  emd_15480_validation.cif.gz emd_15480_validation.cif.gz | 7.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15480 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15480 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15480 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15480 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15480.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15480.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 4.25 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_15480_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
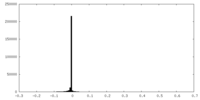
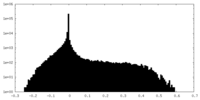
| Density Histograms |
-Half map: #2
| File | emd_15480_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
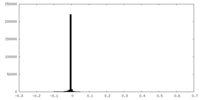
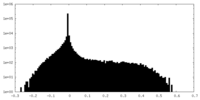
| Density Histograms |
- Sample components
Sample components
-Entire : nucleosome
| Entire | Name: nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: nucleosome
| Supramolecule | Name: nucleosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 210 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: C-flat-2/2 / Material: COPPER/PALLADIUM / Mesh: 200 / Support film - Material: CARBON |
| Vitrification | Cryogen name: OTHER / Details: high pressure freezer HPM010 (ABRA Fluid AG). |
| Details | Drosophila melanogaster embryos were high-pressure frozen. Vitreous sections were cut with a nominal thickness of 75 nm and collected onto 200 mesh C-flat grids |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 2.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.25 µm / Nominal defocus min: 0.25 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 13.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number subtomograms used: 552 |
|---|---|
| Extraction | Number tomograms: 1 / Number images used: 552 |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X