[English] 日本語
 Yorodumi
Yorodumi- EMDB-15213: Cryo-EM density map of Tn4430 TnpA hyperactive mutant (TnpA3X) in... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM density map of Tn4430 TnpA hyperactive mutant (TnpA3X) in complex with IR48 substrate. | |||||||||
 Map data Map data | final auto-sharpened map from Non-Uniform refinement in cryoSPARC v3.3.2 220518 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Tranposition / complex / hyperactive mutant / paired-end complex / DNA BINDING PROTEIN | |||||||||
| Biological species |  | |||||||||
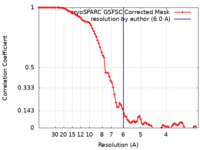
| Method | single particle reconstruction / cryo EM / Resolution: 6.0 Å | |||||||||
 Authors Authors | Shkumatov AV / Liu Y / Efremov RG | |||||||||
| Funding support |  Belgium, 1 items Belgium, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: AFM-based force spectroscopy unravels stepwise formation of the DNA transposition complex in the widespread Tn3 family mobile genetic elements. Authors: Maricruz Fernandez / Alexander V Shkumatov / Yun Liu / Claire Stulemeijer / Sylvie Derclaye / Rouslan G Efremov / Bernard Hallet / David Alsteens /  Abstract: Transposon Tn4430 belongs to a widespread family of bacterial transposons, the Tn3 family, which plays a prevalent role in the dissemination of antibiotic resistance among pathogens. Despite recent ...Transposon Tn4430 belongs to a widespread family of bacterial transposons, the Tn3 family, which plays a prevalent role in the dissemination of antibiotic resistance among pathogens. Despite recent data on the structural architecture of the transposition complex, the molecular mechanisms underlying the replicative transposition of these elements are still poorly understood. Here, we use force-distance curve-based atomic force microscopy to probe the binding of the TnpA transposase of Tn4430 to DNA molecules containing one or two transposon ends and to extract the thermodynamic and kinetic parameters of transposition complex assembly. Comparing wild-type TnpA with previously isolated deregulated TnpA mutants supports a stepwise pathway for transposition complex formation and activation during which TnpA first binds as a dimer to a single transposon end and then undergoes a structural transition that enables it to bind the second end cooperatively and to become activated for transposition catalysis, the latter step occurring at a much faster rate for the TnpA mutants. Our study thus provides an unprecedented approach to probe the dynamic of a complex DNA processing machinery at the single-particle level. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15213.map.gz emd_15213.map.gz | 56.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15213-v30.xml emd-15213-v30.xml emd-15213.xml emd-15213.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15213_fsc.xml emd_15213_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_15213.png emd_15213.png | 116.7 KB | ||
| Masks |  emd_15213_msk_1.map emd_15213_msk_1.map | 59.6 MB |  Mask map Mask map | |
| Others |  emd_15213_half_map_1.map.gz emd_15213_half_map_1.map.gz emd_15213_half_map_2.map.gz emd_15213_half_map_2.map.gz | 55.2 MB 55.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15213 http://ftp.pdbj.org/pub/emdb/structures/EMD-15213 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15213 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15213 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15213.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15213.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | final auto-sharpened map from Non-Uniform refinement in cryoSPARC v3.3.2 220518 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.568 Å | ||||||||||||||||||||||||||||||||||||
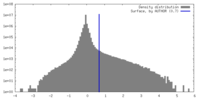
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15213_msk_1.map emd_15213_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_15213_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_15213_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TnpA3X-IR48 complex
| Entire | Name: TnpA3X-IR48 complex |
|---|---|
| Components |
|
-Supramolecule #1: TnpA3X-IR48 complex
| Supramolecule | Name: TnpA3X-IR48 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: hyperactive TnpA mutant (W24R, A174V, E740G) in complex with 48 bp DNA containing transposon recognition sequence. |
|---|
-Supramolecule #2: TnpA 3X mutant
| Supramolecule | Name: TnpA 3X mutant / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 / Details: hyperactive TnpA mutant (W24R, A174V, E740G) |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: IR48 substrate
| Supramolecule | Name: IR48 substrate / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 / Details: DNA substrate |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transposase for transposon Tn4430
| Macromolecule | Name: Transposase for transposon Tn4430 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MGVKQLLSEA QRNELMDLSR LTERDLVTFH TFSKHDLHLI LKHRRGYNRL GFALQLVLIR YPGWSLTEYK DIPQYVVAYV ASQLQIPPE EFLVYAKRGN TLWEHLGEIR TEYGYQNFSS EYKETLLQFL VQQAMDNNNT LYLIEITIST LRKMKVILPA M YVIEDIVW ...String: MGVKQLLSEA QRNELMDLSR LTERDLVTFH TFSKHDLHLI LKHRRGYNRL GFALQLVLIR YPGWSLTEYK DIPQYVVAYV ASQLQIPPE EFLVYAKRGN TLWEHLGEIR TEYGYQNFSS EYKETLLQFL VQQAMDNNNT LYLIEITIST LRKMKVILPA M YVIEDIVW EAKQQVDQKV YSILHDGLVQ EQKDQLDALL LPTINGKSPL AWLKDVPAQP SPESFLKVID RLQFVQKIGL TI DTTKINT NRLRQLARLG SKYEPYAFRR FNEVKRYSML VSFLLEITQD LIDYAIEIHD RLMMNLQTKG KKEQDEIQQA NGK KLNEKI LQFITVCGTL IEAKETGKDA FAALDEVMSW NEMVESVEEA KQLSRPLNYD YLDLLNTRYS YVRRYAPTLL RSLH FRATK SGEPVLQALD TIHELNETGK RKVPHGAPLH FVSNRWQKHV YDDDGNINRH YYELAALTEL RNHIRSGDIF VSGSR HHKA FDDYLIPYDE WNEVSNIPNG LTAPLKAEDY ITDRINRLNE HLEWLSKNSE KLEGVDISQG KLHVERLDRG TPEEAK AFS KLLHSMLPRI KLTDLLIEVA SWTGFHDQFI HASTNQSPDQ EEQNIVLATL MAMGTNIGLT KMAEATPGIS YRQMANA SQ WRMYDDAMVR AQSILVNFQK EQKLSSYWGD GTTSSSDGMR LSIAVRSLHA DSNPHYGTGK GGTIYRFVSD QLSAYHVK V ITTNARDALH VLDGLLHHGT DLKIEEHYTD TAGYTDQVFA LTHLLGFRFA PRIRDLADTK LFSIPGGEEY ENVQALLKG KINVKLIKEN YEDIRRLAYS VQTGKVSSAL IMGKLGSYAR QNKLATALGE MGRIEKTLFT LDYISNKAVR RRVQKGLNKG EAINALARI IFFGQRGEFR ERALQDQLQR ASALNIIINA ISVWNTVYME KAVEELKARG EFREDLMPYA WPLGWEHINF L GEYKFEGL HDTGQMNLRP LRIKEPFYSP IRSFLEQKLI SEEDLNSAVD HHHHHH |
-Macromolecule #2: IR48 DNA substrate, non transferred strand
| Macromolecule | Name: IR48 DNA substrate, non transferred strand / type: dna / ID: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: (DC)(DC)(DA)(DT)(DG)(DG)(DG)(DG)(DG)(DT) (DA)(DC)(DC)(DG)(DC)(DC)(DA)(DG)(DC)(DA) (DT)(DT)(DT)(DC)(DG)(DG)(DA)(DA)(DA) (DA)(DA)(DA)(DA)(DC)(DC)(DA)(DC)(DG)(DC) (DT) (DA)(DA)(DG)(DA)(DT)(DC)(DC)(DT) |
-Macromolecule #3: IR48 transferred strand
| Macromolecule | Name: IR48 transferred strand / type: dna / ID: 3 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: (DA)(DG)(DG)(DA)(DT)(DC)(DT)(DT)(DA)(DG) (DC)(DG)(DT)(DG)(DG)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DC)(DC)(DG)(DA)(DA)(DA)(DT) (DG)(DC)(DT)(DG)(DG)(DC)(DG)(DG)(DT)(DA) (DC) (DC)(DC)(DC)(DC)(DA)(DT)(DG)(DG) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 50 mM HEPES (pH 7.5), 100 mM NaCl, 30 mM L-Arg HCL |
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 0.035 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: GATAN CRYOPLUNGE 3 |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 6734 / Average exposure time: 3.0 sec. / Average electron dose: 64.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)