+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of Connexin 32 gap junction channel | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | connexin / Hemi channel / cell communication / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationOligomerization of connexins into connexons / Transport of connexins along the secretory pathway / gap junction assembly / connexin complex / Gap junction assembly / gap junction channel activity / cell-cell signaling / nervous system development / endoplasmic reticulum membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
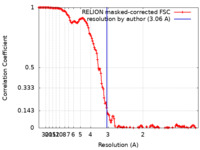
| Method | single particle reconstruction / cryo EM / Resolution: 3.06 Å | |||||||||
 Authors Authors | Qi C / Korkhov VM | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Structures of wild-type and selected CMT1X mutant connexin 32 gap junction channels and hemichannels. Authors: Chao Qi / Pia Lavriha / Erva Bayraktar / Anand Vaithia / Dina Schuster / Micaela Pannella / Valentina Sala / Paola Picotti / Mario Bortolozzi / Volodymyr M Korkhov /   Abstract: In myelinating Schwann cells, connection between myelin layers is mediated by gap junction channels (GJCs) formed by docked connexin 32 (Cx32) hemichannels (HCs). Mutations in Cx32 cause the X-linked ...In myelinating Schwann cells, connection between myelin layers is mediated by gap junction channels (GJCs) formed by docked connexin 32 (Cx32) hemichannels (HCs). Mutations in Cx32 cause the X-linked Charcot-Marie-Tooth disease (CMT1X), a degenerative neuropathy without a cure. A molecular link between Cx32 dysfunction and CMT1X pathogenesis is still missing. Here, we describe the high-resolution cryo-electron cryo-myography (cryo-EM) structures of the Cx32 GJC and HC, along with two CMT1X-linked mutants, W3S and R22G. While the structures of wild-type and mutant GJCs are virtually identical, the HCs show a major difference: In the W3S and R22G mutant HCs, the amino-terminal gating helix partially occludes the pore, consistent with a diminished HC activity. Our results suggest that HC dysfunction may be involved in the pathogenesis of CMT1X. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15011.map.gz emd_15011.map.gz | 21.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15011-v30.xml emd-15011-v30.xml emd-15011.xml emd-15011.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15011_fsc.xml emd_15011_fsc.xml | 13.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_15011.png emd_15011.png | 86.5 KB | ||
| Masks |  emd_15011_msk_1.map emd_15011_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15011.cif.gz emd-15011.cif.gz | 6.1 KB | ||
| Others |  emd_15011_half_map_1.map.gz emd_15011_half_map_1.map.gz emd_15011_half_map_2.map.gz emd_15011_half_map_2.map.gz | 202.3 MB 202 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15011 http://ftp.pdbj.org/pub/emdb/structures/EMD-15011 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15011 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15011 | HTTPS FTP |
-Related structure data
| Related structure data |  7zxnMC  7zxmC  7zxoC  7zxpC  7zxqC  7zxtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15011.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15011.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.654 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15011_msk_1.map emd_15011_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
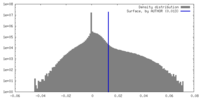
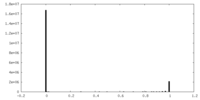
| Density Histograms |
-Half map: #1
| File | emd_15011_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
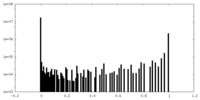
| Density Histograms |
-Half map: #2
| File | emd_15011_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
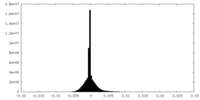
| Density Histograms |
- Sample components
Sample components
-Entire : Connexin32 gap junction channel complex
| Entire | Name: Connexin32 gap junction channel complex |
|---|---|
| Components |
|
-Supramolecule #1: Connexin32 gap junction channel complex
| Supramolecule | Name: Connexin32 gap junction channel complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32 kDa/nm |
-Macromolecule #1: Gap junction beta-1 protein
| Macromolecule | Name: Gap junction beta-1 protein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.065533 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MNWTGLYTLL SGVNRHSTAI GRVWLSVIFI FRIMVLVVAA ESVWGDEKSS FICNTLQPGC NSVCYDQFFP ISHVRLWSLQ LILVSTPAL LVAMHVAHQQ HIEKKMLRLE GHGDPLHLEE VKRHKVHISG TLWWTYVISV VFRLLFEAVF MYVFYLLYPG Y AMVRLVKC ...String: MNWTGLYTLL SGVNRHSTAI GRVWLSVIFI FRIMVLVVAA ESVWGDEKSS FICNTLQPGC NSVCYDQFFP ISHVRLWSLQ LILVSTPAL LVAMHVAHQQ HIEKKMLRLE GHGDPLHLEE VKRHKVHISG TLWWTYVISV VFRLLFEAVF MYVFYLLYPG Y AMVRLVKC DVYPCPNTVD CFVSRPTEKT VFTVFMLAAS GICIILNVAE VVYLIIRACA RRAQRRSNPP SRKGSGFGHR LS PEYKQNE INKLLSEQDG SLKDILRRSP GTGAGLAEKS DRCSAC UniProtKB: Gap junction beta-1 protein |
-Macromolecule #2: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 2 / Number of copies: 6 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)