+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | ARISC-RAP80 complex, Map 5 | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.9 Å | |||||||||||||||
 Authors Authors | Foglizzo M / Zeqiraj E | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: J Cell Biol / Year: 2022 Journal: J Cell Biol / Year: 2022Title: Autologous K63 deubiquitylation within the BRCA1-A complex licenses DNA damage recognition. Authors: Qinqin Jiang / Martina Foglizzo / Yaroslav I Morozov / Xuejiao Yang / Arindam Datta / Lei Tian / Vaughn Thada / Weihua Li / Elton Zeqiraj / Roger A Greenberg /   Abstract: The BRCA1-A complex contains matching lysine-63 ubiquitin (K63-Ub) binding and deubiquitylating activities. How these functionalities are coordinated to effectively respond to DNA damage remains ...The BRCA1-A complex contains matching lysine-63 ubiquitin (K63-Ub) binding and deubiquitylating activities. How these functionalities are coordinated to effectively respond to DNA damage remains unknown. We generated Brcc36 deubiquitylating enzyme (DUB) inactive mice to address this gap in knowledge in a physiologic system. DUB inactivation impaired BRCA1-A complex damage localization and repair activities while causing early lethality when combined with Brca2 mutation. Damage response dysfunction in DUB-inactive cells corresponded to increased K63-Ub on RAP80 and BRCC36. Chemical cross-linking coupled with liquid chromatography-tandem mass spectrometry (LC-MS/MS) and cryogenic-electron microscopy (cryo-EM) analyses of isolated BRCA1-A complexes demonstrated the RAP80 ubiquitin interaction motifs are occupied by ubiquitin exclusively in the DUB-inactive complex, linking auto-inhibition by internal K63-Ub chains to loss of damage site ubiquitin recognition. These findings identify RAP80 and BRCC36 as autologous DUB substrates in the BRCA1-A complex, thus explaining the evolution of matching ubiquitin-binding and hydrolysis activities within a single macromolecular assembly. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15003.map.gz emd_15003.map.gz | 74.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15003-v30.xml emd-15003-v30.xml emd-15003.xml emd-15003.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15003.png emd_15003.png | 32.9 KB | ||
| Masks |  emd_15003_msk_1.map emd_15003_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_15003_half_map_1.map.gz emd_15003_half_map_1.map.gz emd_15003_half_map_2.map.gz emd_15003_half_map_2.map.gz | 98.2 MB 98.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15003 http://ftp.pdbj.org/pub/emdb/structures/EMD-15003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15003 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15003.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15003.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.065 Å | ||||||||||||||||||||||||||||||||||||
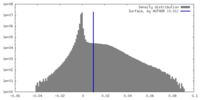
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15003_msk_1.map emd_15003_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
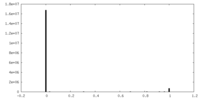
| Density Histograms |
-Half map: #1
| File | emd_15003_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
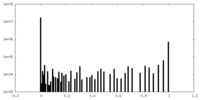
| Density Histograms |
-Half map: #2
| File | emd_15003_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
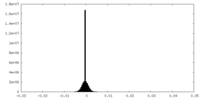
| Density Histograms |
- Sample components
Sample components
-Entire : Complex containing the four-subunit ARISC together with FL RAP80
| Entire | Name: Complex containing the four-subunit ARISC together with FL RAP80 |
|---|---|
| Components |
|
-Supramolecule #1: Complex containing the four-subunit ARISC together with FL RAP80
| Supramolecule | Name: Complex containing the four-subunit ARISC together with FL RAP80 type: complex / Chimera: Yes / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Molecular weight | Experimental: 235 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.16 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: The sample was supplemented with 2 mM EDTA/EGTA mixture immediately before grids preparation | ||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: The sample was blotted for 6 seconds and with a blot force of 6 before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4392 / Average exposure time: 70.0 sec. / Average electron dose: 1.46 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.1 µm / Nominal defocus min: 1.7 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)