[English] 日本語
 Yorodumi
Yorodumi- EMDB-14675: IF(heme/confined) conformation of CydDC mutant (E500Q.C) in AMP-P... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | IF(heme/confined) conformation of CydDC mutant (E500Q.C) in AMP-PNP(CydD) bound state (Dataset-22) | |||||||||
 Map data Map data | RELION local-resolution filtered map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ABC transporter / CydDC / Heme transporter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcysteine export across plasma membrane / ATP-binding cassette (ABC) transporter complex, integrated substrate binding / cytochrome biosynthetic process / Translocases; Catalysing the translocation of amino acids and peptides; Linked to the hydrolysis of a nucleoside triphosphate / ABC-type heme transporter activity / glutathione transmembrane transport / heme transmembrane transport / ATPase-coupled lipid transmembrane transporter activity / ATPase-coupled transmembrane transporter activity / ATP-binding cassette (ABC) transporter complex ...cysteine export across plasma membrane / ATP-binding cassette (ABC) transporter complex, integrated substrate binding / cytochrome biosynthetic process / Translocases; Catalysing the translocation of amino acids and peptides; Linked to the hydrolysis of a nucleoside triphosphate / ABC-type heme transporter activity / glutathione transmembrane transport / heme transmembrane transport / ATPase-coupled lipid transmembrane transporter activity / ATPase-coupled transmembrane transporter activity / ATP-binding cassette (ABC) transporter complex / cell redox homeostasis / transmembrane transport / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
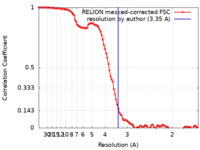
| Method | single particle reconstruction / cryo EM / Resolution: 3.35 Å | |||||||||
 Authors Authors | Wu D / Safarian S | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2023 Journal: Nat Chem Biol / Year: 2023Title: Dissecting the conformational complexity and mechanism of a bacterial heme transporter. Authors: Di Wu / Ahmad R Mehdipour / Franziska Finke / Hojjat G Goojani / Roan R Groh / Tamara N Grund / Thomas M B Reichhart / Rita Zimmermann / Sonja Welsch / Dirk Bald / Mark Shepherd / Gerhard ...Authors: Di Wu / Ahmad R Mehdipour / Franziska Finke / Hojjat G Goojani / Roan R Groh / Tamara N Grund / Thomas M B Reichhart / Rita Zimmermann / Sonja Welsch / Dirk Bald / Mark Shepherd / Gerhard Hummer / Schara Safarian /      Abstract: Iron-bound cyclic tetrapyrroles (hemes) are redox-active cofactors in bioenergetic enzymes. However, the mechanisms of heme transport and insertion into respiratory chain complexes remain unclear. ...Iron-bound cyclic tetrapyrroles (hemes) are redox-active cofactors in bioenergetic enzymes. However, the mechanisms of heme transport and insertion into respiratory chain complexes remain unclear. Here, we used cellular, biochemical, structural and computational methods to characterize the structure and function of the heterodimeric bacterial ABC transporter CydDC. We provide multi-level evidence that CydDC is a heme transporter required for functional maturation of cytochrome bd, a pharmaceutically relevant drug target. Our systematic single-particle cryogenic-electron microscopy approach combined with atomistic molecular dynamics simulations provides detailed insight into the conformational landscape of CydDC during substrate binding and occlusion. Our simulations reveal that heme binds laterally from the membrane space to the transmembrane region of CydDC, enabled by a highly asymmetrical inward-facing CydDC conformation. During the binding process, heme propionates interact with positively charged residues on the surface and later in the substrate-binding pocket of the transporter, causing the heme orientation to rotate 180°. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14675.map.gz emd_14675.map.gz | 56.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14675-v30.xml emd-14675-v30.xml emd-14675.xml emd-14675.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14675_fsc.xml emd_14675_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_14675.png emd_14675.png | 77.3 KB | ||
| Filedesc metadata |  emd-14675.cif.gz emd-14675.cif.gz | 6.5 KB | ||
| Others |  emd_14675_additional_1.map.gz emd_14675_additional_1.map.gz emd_14675_additional_2.map.gz emd_14675_additional_2.map.gz emd_14675_half_map_1.map.gz emd_14675_half_map_1.map.gz emd_14675_half_map_2.map.gz emd_14675_half_map_2.map.gz | 7.7 MB 71.2 MB 71.3 MB 71.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14675 http://ftp.pdbj.org/pub/emdb/structures/EMD-14675 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14675 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14675 | HTTPS FTP |
-Validation report
| Summary document |  emd_14675_validation.pdf.gz emd_14675_validation.pdf.gz | 910.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14675_full_validation.pdf.gz emd_14675_full_validation.pdf.gz | 910 KB | Display | |
| Data in XML |  emd_14675_validation.xml.gz emd_14675_validation.xml.gz | 17.6 KB | Display | |
| Data in CIF |  emd_14675_validation.cif.gz emd_14675_validation.cif.gz | 23.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14675 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14675 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14675 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14675 | HTTPS FTP |
-Related structure data
| Related structure data |  7zdwMC  7zd5C  7zdaC  7zdbC  7zdcC  7zdeC  7zdfC  7zdgC  7zdkC  7zdlC  7zdrC  7zdsC  7zdtC  7zduC  7zdvC  7ze5C  7zecC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14675.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14675.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION local-resolution filtered map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: RELION masked postprocessing map
| File | emd_14675_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION masked postprocessing map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: RELION masked 3D refinement map
| File | emd_14675_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION masked 3D refinement map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: RELION 3D refinement half map 1
| File | emd_14675_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION 3D refinement half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: RELION 3D refinement half map 2
| File | emd_14675_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION 3D refinement half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CydDC heterodimer
| Entire | Name: CydDC heterodimer |
|---|---|
| Components |
|
-Supramolecule #1: CydDC heterodimer
| Supramolecule | Name: CydDC heterodimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 130 KDa |
-Macromolecule #1: ATP-binding/permease protein CydC
| Macromolecule | Name: ATP-binding/permease protein CydC / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 62.979867 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRALLPYLAL YKRHKWMLSL GIVLAIVTLL ASIGLLTLSG WFLSASAVAG VAGLYSFNYM LPAAGVRGAA ITRTAGRYFE RLVSHDATF RVLQHLRIYT FSKLLPLSPA GLARYRQGEL LNRVVADVDT LDHLYLRVIS PLVGAFVVIM VVTIGLSFLD F TLAFTLGG ...String: MRALLPYLAL YKRHKWMLSL GIVLAIVTLL ASIGLLTLSG WFLSASAVAG VAGLYSFNYM LPAAGVRGAA ITRTAGRYFE RLVSHDATF RVLQHLRIYT FSKLLPLSPA GLARYRQGEL LNRVVADVDT LDHLYLRVIS PLVGAFVVIM VVTIGLSFLD F TLAFTLGG IMLLTLFLMP PLFYRAGKST GQNLTHLRGQ YRQQLTAWLQ GQAELTIFGA SDRYRTQLEN TEIQWLEAQR RQ SELTALS QAIMLLIGAL AVILMLWMAS GGVGGNAQPG ALIALFVFCA LAAFEALAPV TGAFQHLGQV IASAVRISDL TDQ KPEVTF PDTQTRVADR VSLTLRDVQF TYPEQSQQAL KGISLQVNAG EHIAILGRTG CGKSTLLQQL TRAWDPQQGE ILLN DSPIA SLNEAALRQT ISVVPQRVHL FSATLRDNLL LASPGSSDEA LSEILRRVGL EKLLEDAGLN SWLGEGGRQL SGGEL RRLA IARALLHDAP LVLLDQPTEG LDATTESQIL ELLAEMMREK TVLMVTHRLR GLSRFQQIIV MDNGQIIEQG THAELL ARQ GRYYQFKQGL UniProtKB: Glutathione/L-cysteine transport system ATP-binding/permease protein CydC |
-Macromolecule #2: ATP-binding/permease protein CydD
| Macromolecule | Name: ATP-binding/permease protein CydD / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 65.118867 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNKSRQKELT RWLKQQSVIS QRWLNISRLL GFVSGILIIA QAWFMARILQ HMIMENIPRE ALLLPFTLLV LTFVLRAWVV WLRERVGYH AGQHIRFAIR RQVLDRLQQA GPAWIQGKPA GSWATLVLEQ IDDMHDYYAR YLPQMALAVS VPLLIVVAIF P SNWAAALI ...String: MNKSRQKELT RWLKQQSVIS QRWLNISRLL GFVSGILIIA QAWFMARILQ HMIMENIPRE ALLLPFTLLV LTFVLRAWVV WLRERVGYH AGQHIRFAIR RQVLDRLQQA GPAWIQGKPA GSWATLVLEQ IDDMHDYYAR YLPQMALAVS VPLLIVVAIF P SNWAAALI LLGTAPLIPL FMALVGMGAA DANRRNFLAL ARLSGHFLDR LRGMETLRIF GRGEAEIESI RSASEDFRQR TM EVLRLAF LSSGILEFFT SLSIALVAVY FGFSYLGELD FGHYDTGVTL AAGFLALILA PEFFQPLRDL GTFYHAKAQA VGA ADSLKT FMETPLAHPQ RGEAELASTD PVTIEAEELF ITSPEGKTLA GPLNFTLPAG QRAVLVGRSG SGKSSLLNAL SGFL SYQGS LRINGIELRD LSPESWRKHL SWVGQNPQLP AATLRDNVLL ARPDASEQEL QAALDNAWVS EFLPLLPQGV DTPVG DQAA RLSVGQAQRV AVARALLNPC SLLLLDEPAA SLDAHSEQRV MEALNAASLR QTTLMVTHQL EDLADWDVIW VMQDGR IIE QGRYAELSVA GGPFATLLAH RQEEI UniProtKB: Glutathione/L-cysteine transport system ATP-binding/permease protein CydD |
-Macromolecule #3: HEME B/C
| Macromolecule | Name: HEME B/C / type: ligand / ID: 3 / Number of copies: 1 / Formula: HEB |
|---|---|
| Molecular weight | Theoretical: 618.503 Da |
| Chemical component information |  ChemComp-HEB: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / type: ligand / ID: 5 / Number of copies: 1 / Formula: ANP |
|---|---|
| Molecular weight | Theoretical: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 6 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.1 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)