+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 | ||||||||||||
 Map data Map data | AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (main map) | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
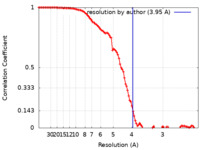
| Method | single particle reconstruction / cryo EM / Resolution: 3.95 Å | ||||||||||||
 Authors Authors | van Schooten J / Ward A | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Identification of IOMA-class neutralizing antibodies targeting the CD4-binding site on the HIV-1 envelope glycoprotein. Authors: Jelle van Schooten / Elinaz Farokhi / Anna Schorcht / Tom L G M van den Kerkhof / Hongmei Gao / Patricia van der Woude / Judith A Burger / Tim G Rijkhold Meesters / Tom Bijl / Riham ...Authors: Jelle van Schooten / Elinaz Farokhi / Anna Schorcht / Tom L G M van den Kerkhof / Hongmei Gao / Patricia van der Woude / Judith A Burger / Tim G Rijkhold Meesters / Tom Bijl / Riham Ghalaiyini / Hannah L Turner / Jessica Dorning / Barbera D C van Schaik / Antoine H C van Kampen / Celia C Labranche / Robyn L Stanfield / Devin Sok / David C Montefiori / Dennis R Burton / Michael S Seaman / Gabriel Ozorowski / Ian A Wilson / Rogier W Sanders / Andrew B Ward / Marit J van Gils /   Abstract: A major goal of current HIV-1 vaccine design efforts is to induce broadly neutralizing antibodies (bNAbs). The VH1-2-derived bNAb IOMA directed to the CD4-binding site of the HIV-1 envelope ...A major goal of current HIV-1 vaccine design efforts is to induce broadly neutralizing antibodies (bNAbs). The VH1-2-derived bNAb IOMA directed to the CD4-binding site of the HIV-1 envelope glycoprotein is of interest because, unlike the better-known VH1-2-derived VRC01-class bNAbs, it does not require a rare short light chain complementarity-determining region 3 (CDRL3). Here, we describe three IOMA-class NAbs, ACS101-103, with up to 37% breadth, that share many characteristics with IOMA, including an average-length CDRL3. Cryo-electron microscopy revealed that ACS101 shares interactions with those observed with other VH1-2 and VH1-46-class bNAbs, but exhibits a unique binding mode to residues in loop D. Analysis of longitudinal sequences from the patient suggests that a transmitter/founder-virus lacking the N276 glycan might have initiated the development of these NAbs. Together these data strengthen the rationale for germline-targeting vaccination strategies to induce IOMA-class bNAbs and provide a wealth of sequence and structural information to support such strategies. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14474.map.gz emd_14474.map.gz | 97.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14474-v30.xml emd-14474-v30.xml emd-14474.xml emd-14474.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14474_fsc.xml emd_14474_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_14474.png emd_14474.png | 91.5 KB | ||
| Masks |  emd_14474_msk_1.map emd_14474_msk_1.map | 103 MB |  Mask map Mask map | |
| Others |  emd_14474_additional_1.map.gz emd_14474_additional_1.map.gz emd_14474_additional_2.map.gz emd_14474_additional_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14474 http://ftp.pdbj.org/pub/emdb/structures/EMD-14474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14474 | HTTPS FTP |
-Validation report
| Summary document |  emd_14474_validation.pdf.gz emd_14474_validation.pdf.gz | 482.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14474_full_validation.pdf.gz emd_14474_full_validation.pdf.gz | 482.4 KB | Display | |
| Data in XML |  emd_14474_validation.xml.gz emd_14474_validation.xml.gz | 11.7 KB | Display | |
| Data in CIF |  emd_14474_validation.cif.gz emd_14474_validation.cif.gz | 15.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14474 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14474 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14474 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14474 | HTTPS FTP |
-Related structure data
| Related structure data |  7z3aMC  7u04C  7u0kC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14474.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14474.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (main map) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||
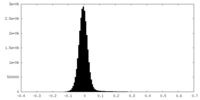
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14474_msk_1.map emd_14474_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
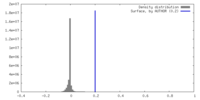
| Density Histograms |
-Additional map: AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (main map)
| File | emd_14474_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (main map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
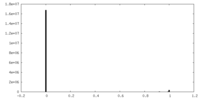
| Density Histograms |
-Additional map: AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (half map A)
| File | emd_14474_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124 (half map A) | ||||||||||||
| Projections & Slices |
| ||||||||||||
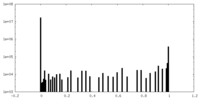
| Density Histograms |
- Sample components
Sample components
+Entire : AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124
+Supramolecule #1: AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124
+Supramolecule #2: AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124
+Supramolecule #3: AMC009 SOSIPv5.2 in complex with Fabs ACS101 and ACS124
+Macromolecule #1: AMC009 SOSIPv5.2 envelope glycoprotein gp120
+Macromolecule #2: AMC009 SOSIP.v5.2 envelope glycoprotein gp41
+Macromolecule #3: ACS124 heavy chain
+Macromolecule #4: ACS124 light chain
+Macromolecule #5: ACS101 heavy chain
+Macromolecule #6: ACS101 light chain
+Macromolecule #11: 2-acetamido-2-deoxy-beta-D-glucopyranose
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)