[English] 日本語
 Yorodumi
Yorodumi- EMDB-14378: Cryo-EM structure of Mycobacterium tuberculosis RNA polymerase ho... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Mycobacterium tuberculosis RNA polymerase holoenzyme dimer comprising sigma factor SigB | |||||||||
 Map data Map data | RNA polymerase dimer Class 1 (conformation 1) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) | |||||||||
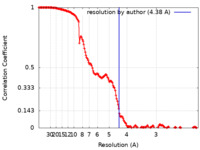
| Method | single particle reconstruction / cryo EM / Resolution: 4.38 Å | |||||||||
 Authors Authors | Brodolin K | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation | Journal: Nucleic Acids Res / Year: 2014 Title: Mycobacterium RbpA cooperates with the stress-response σB subunit of RNA polymerase in promoter DNA unwinding. Authors: Yangbo Hu / Zakia Morichaud / Ayyappasamy Sudalaiyadum Perumal / Françoise Roquet-Baneres / Konstantin Brodolin /  Abstract: RbpA, a transcriptional activator that is essential for Mycobacterium tuberculosis replication and survival during antibiotic treatment, binds to RNA polymerase (RNAP) in the absence of promoter DNA. ...RbpA, a transcriptional activator that is essential for Mycobacterium tuberculosis replication and survival during antibiotic treatment, binds to RNA polymerase (RNAP) in the absence of promoter DNA. It has been hypothesized that RbpA stimulates housekeeping gene expression by promoting assembly of the σ(A) subunit with core RNAP. Here, using a purified in vitro transcription system of M. tuberculosis, we show that RbpA functions in a promoter-dependent manner as a companion of RNAP essential for promoter DNA unwinding and formation of the catalytically active open promoter complex (RPo). Screening for RbpA activity using a full panel of the M. tuberculosis σ subunits demonstrated that RbpA targets σ(A) and stress-response σ(B), but not the alternative σ subunits from the groups 3 and 4. In contrast to σ(A), the σ(B) subunit activity displayed stringent dependency upon RbpA. These results suggest that RbpA-dependent control of RPo formation provides a mechanism for tuning gene expression during the switch between different physiological states, and in the stress response. #1:  Journal: Biorxiv / Year: 2022 Journal: Biorxiv / Year: 2022Title: Structural basis of the mycobacterial stress-response RNA polymerase auto-inhibition via oligomerization Authors: Morichaud Z / Trapani S / Vishwakarma R / Chaloin L / Lionne C / Lai-Kee-Him J / Bron P / Brodolin K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14378.map.gz emd_14378.map.gz | 327.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14378-v30.xml emd-14378-v30.xml emd-14378.xml emd-14378.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14378_fsc.xml emd_14378_fsc.xml | 15.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_14378.png emd_14378.png | 138.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14378 http://ftp.pdbj.org/pub/emdb/structures/EMD-14378 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14378 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14378 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14378.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14378.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RNA polymerase dimer Class 1 (conformation 1) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : RNA polymerase holoenzyme with sigma factor sigB
| Entire | Name: RNA polymerase holoenzyme with sigma factor sigB |
|---|---|
| Components |
|
-Supramolecule #1: RNA polymerase holoenzyme with sigma factor sigB
| Supramolecule | Name: RNA polymerase holoenzyme with sigma factor sigB / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all Details: RNA polymerase holoenzyme assembled from individually expressed RNA polymerase core and sigma factor sigB |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Molecular weight | Theoretical: 800 KDa |
-Macromolecule #1: DNA-directed RNA polymerase subunit alpha
| Macromolecule | Name: DNA-directed RNA polymerase subunit alpha / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: MLISQRPTLS EDVLTDNRSQ FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTEIILNL KSLVVSSEED EPVTMYLRKQ GPGEVTAGDI VPPAGVTVHN PGMHIATLND KGKLEVELVV ERGRGYVPAV QNRASGAEIG RIPVDSIYSP ...String: MLISQRPTLS EDVLTDNRSQ FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTEIILNL KSLVVSSEED EPVTMYLRKQ GPGEVTAGDI VPPAGVTVHN PGMHIATLND KGKLEVELVV ERGRGYVPAV QNRASGAEIG RIPVDSIYSP VLKVTYKVDA TRVEQRTDFD KLILDVETKN SISPRDALAS AGKTLVELFG LARELNVEAE GIEIGPSPAE ADHIASFALP IDDLDLTVRS YNCLKREGVH TVGELVARTE SDLLDIRNFG QKSIDEVKIK LHQLGLSLKD SPPSFDPSEV AGYDVATGTW STEGAYDEQD YAETEQL |
-Macromolecule #2: DNA-directed RNA polymerase subunit alpha
| Macromolecule | Name: DNA-directed RNA polymerase subunit alpha / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: MLISQRPTLS EDVLTDNRSQ FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTEIILNL KSLVVSSEED EPVTMYLRKQ GPGEVTAGDI VPPAGVTVHN PGMHIATLND KGKLEVELVV ERGRGYVPAV QNRASGAEIG RIPVDSIYSP ...String: MLISQRPTLS EDVLTDNRSQ FVIEPLEPGF GYTLGNSLRR TLLSSIPGAA VTSIRIDGVL HEFTTVPGVK EDVTEIILNL KSLVVSSEED EPVTMYLRKQ GPGEVTAGDI VPPAGVTVHN PGMHIATLND KGKLEVELVV ERGRGYVPAV QNRASGAEIG RIPVDSIYSP VLKVTYKVDA TRVEQRTDFD KLILDVETKN SISPRDALAS AGKTLVELFG LARELNVEAE GIEIGPSPAE ADHIASFALP IDDLDLTVRS YNCLKREGVH TVGELVARTE SDLLDIRNFG QKSIDEVKIK LHQLGLSLKD SPPSFDPSEV AGYDVATGTW STEGAYDEQD YAETEQL |
-Macromolecule #3: DNA-directed RNA polymerase subunit beta
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta / type: protein_or_peptide / ID: 3 / Details: L2E3G4C5 -> V / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: MVLADSRQSK TAASPSPSRP QSSSNNSVPG APNRVSFAKL REPLEVPGLL DVQTDSFEWL IGSPRWRESA AERGDVNPVG GLEEVLYELS PIEDFSGSMS LSFSDPRFDD VKAPVDECKD KDMTYAAPLF VTAEFINNNT GEIKSQTVFM GDFPMMTEKG TFIINGTERV ...String: MVLADSRQSK TAASPSPSRP QSSSNNSVPG APNRVSFAKL REPLEVPGLL DVQTDSFEWL IGSPRWRESA AERGDVNPVG GLEEVLYELS PIEDFSGSMS LSFSDPRFDD VKAPVDECKD KDMTYAAPLF VTAEFINNNT GEIKSQTVFM GDFPMMTEKG TFIINGTERV VVSQLVRSPG VYFDETIDKS TDKTLHSVKV IPSRGAWLEF DVDKRDTVGV RIDRKRRQPV TVLLKALGWT SEQIVERFGF SEIMRSTLEK DNTVGTDEAL LDIYRKLRPG EPPTKESAQT LLENLFFKEK RYDLARVGRY KVNKKLGLHV GEPITSSTLT EEDVVATIEY LVRLHEGQTT MTVPGGVEVP VETDDIDHFG NRRLRTVGEL IQNQIRVGMS RMERVVRERM TTQDVEAITP QTLINIRPVV AAIKEFFGTS QLSQFMDQNN PLSGLTHKRR LSALGPGGLS RERAGLEVRD VHPSHYGRMC PIETPEGPNI GLIGSLSVYA RVNPFGFIET PYRKVVDGVV SDEIVYLTAD EEDRHVVAQA NSPIDADGRF VEPRVLVRRK AGEVEYVPSS EVDYMDVSPR QMVSVATAMI PFLEHDDANR ALMGANMQRQ AVPLVRSEAP LVGTGMELRA AIDAGDVVVA EESGVIEEVS ADYITVMHDN GTRRTYRMRK FARSNHGTCA NQCPIVDAGD RVEAGQVIAD GPCTDDGEMA LGKNLLVAIM PWEGHNYEDA IILSNRLVEE DVLTSIHIEE HEIDARDTKL GAEEITRDIP NISDEVLADL DERGIVRIGA EVRDGDILVG KVTPKGETEL TPEERLLRAI FGEKAREVRD TSLKVPHGES GKVIGIRVFS REDEDELPAG VNELVRVYVA QKRKISDGDK LAGRHGNKGV IGKILPVEDM PFLADGTPVD IILNTHGVPR RMNIGQILET HLGWCAHSGW KVDAAKGVPD WAARLPDELL EAQPNAIVST PVFDGAQEAE LQGLLSCTLP NRDGDVLVDA DGKAMLFDGR SGEPFPYPVT VGYMYIMKLH HLVDDKIHAR STGPYSMITQ QPLGGKAQFG GQRFGEMECW AMQAYGAAYT LQELLTIKSD DTVGRVKVYE AIVKGENIPE PGIPESFKVL LKELQSLCLN VEVLSSDGAA IELREGEDED LERAAANLGI NLSRNESASV EDLA |
-Macromolecule #4: DNA-directed RNA polymerase subunit beta'
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta' / type: protein_or_peptide / ID: 4 / Details: contains C-terminal 6xHis-tag / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: VNFFDELRIG LATAEDIRQW SYGEVKKPET INYRTLKPEK DGLFCEKIFG PTRDWECYCG KYKRVRFKGI ICERCGVEVT RAKVRRERMG HIELAAPVTH IWYFKGVPSR LGYLLDLAPK DLEKIIYFAA YVITSVDEEM RHNELSTLEA EMAVERKAVE DQRDGELEAR ...String: VNFFDELRIG LATAEDIRQW SYGEVKKPET INYRTLKPEK DGLFCEKIFG PTRDWECYCG KYKRVRFKGI ICERCGVEVT RAKVRRERMG HIELAAPVTH IWYFKGVPSR LGYLLDLAPK DLEKIIYFAA YVITSVDEEM RHNELSTLEA EMAVERKAVE DQRDGELEAR AQKLEADLAE LEAEGAKADA RRKVRDGGER EMRQIRDRAQ RELDRLEDIW STFTKLAPKQ LIVDENLYRE LVDRYGEYFT GAMGAESIQK LIENFDIDAE AESLRDVIRN GKGQKKLRAL KRLKVVAAFQ QSGNSPMGMV LDAVPVIPPE LRPMVQLDGG RFATSDLNDL YRRVINRNNR LKRLIDLGAP EIIVNNEKRM LQESVDALFD NGRRGRPVTG PGNRPLKSLS DLLKGKQGRF RQNLLGKRVD YSGRSVIVVG PQLKLHQCGL PKLMALELFK PFVMKRLVDL NHAQNIKSAK RMVERQRPQV WDVLEEVIAE HPVLLNRAPT LHRLGIQAFE PMLVEGKAIQ LHPLVCEAFN ADFDGDQMAV HLPLSAEAQA EARILMLSSN NILSPASGRP LAMPRLDMVT GLYYLTTEVP GDTGEYQPAS GDHPETGVYS SPAEAIMAAD RGVLSVRAKI KVRLTQLRPP VEIEAELFGH SGWQPGDAWM AETTLGRVMF NELLPLGYPF VNKQMHKKVQ AAIINDLAER YPMIVVAQTV DKLKDAGFYW ATRSGVTVSM ADVLVPPRKK EILDHYEERA DKVEKQFQRG ALNHDERNEA LVEIWKEATD EVGQALREHY PDDNPIITIV DSGATGNFTQ TRTLAGMKGL VTNPKGEFIP RPVKSSFREG LTVLEYFINT HGARKGLADT ALRTADSGYL TRRLVDVSQD VIVREHDCQT ERGIVVELAE RAPDGTLIRD PYIETSAYAR TLGTDAVDEA GNVIVERGQD LGDPEIDALL AAGITQVKVR SVLTCATSTG VCATCYGRSM ATGKLVDIGE AVGIVAAQSI GEPGTQLTMR TFHQGGVGED ITGGLPRVQE LFEARVPRGK APIADVTGRV RLEDGERFYK ITIVPDDGGE EVVYDKISKR QRLRVFKHED GSERVLSDGD HVEVGQQLME GSADPHEVLR VQGPREVQIH LVREVQEVYR AQGVSIHDKH IEVIVRQMLR RVTIIDSGST EFLPGSLIDR AEFEAENRRV VAEGGEPAAG RPVLMGITKA SLATDSWLSA ASFQETTRVL TDAAINCRSD KLNGLKENVI IGKLIPAGTG INRYRNIAVQ PTEEARAAAY TIPSYEDQYY SPDFGAATGA AVPLDDYGYS DYRHHHHHH |
-Macromolecule #5: DNA-directed RNA polymerase subunit omega
| Macromolecule | Name: DNA-directed RNA polymerase subunit omega / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: MSISQSDASL AAVPAVDQFD PSSGASGGYD TPLGITNPPI DELLDRVSSK YALVIYAAKR ARQINDYYNQ LGEGILEYVG PLVEPGLQEK PLSIALREIH ADLLEHTEGE |
-Macromolecule #6: RNA polymerase sigma factor SigB
| Macromolecule | Name: RNA polymerase sigma factor SigB / type: protein_or_peptide / ID: 6 / Details: contains N-terminal 6xHis-tag / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis H37Rv (bacteria) Mycobacterium tuberculosis H37Rv (bacteria) |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MADAPTRATT SRVDSDLDAQ SPAADLVRVY LNGIGKTALL NAAGEVELAK RIEAGLYAEH LLETRKRLGE NRKRDLAAVV RDGEAARRHL LEANLRLVVS LAKRYTGRGM PLLDLIQEGN LGLIRAMEKF DYTKGFKFST YATWWIRQAI ...String: MGSSHHHHHH SSGLVPRGSH MADAPTRATT SRVDSDLDAQ SPAADLVRVY LNGIGKTALL NAAGEVELAK RIEAGLYAEH LLETRKRLGE NRKRDLAAVV RDGEAARRHL LEANLRLVVS LAKRYTGRGM PLLDLIQEGN LGLIRAMEKF DYTKGFKFST YATWWIRQAI TRGMADQSRT IRLPVHLVEQ VNKLARIKRE MHQHLGREAT DEELAAESGI PIDKINDLLE HSRDPVSLDM PVGSEEEAPL GDFIEDAEAM SAENAVIAEL LHTDIRSVLA TLDEREHQVI RLRFGLDDGQ PRTLDQIGKL FGLSRERVRQ IERDVMSKLR HGERADRLRS YAS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.4 mg/mL |
|---|---|
| Buffer | pH: 7.9 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 3064 / Average electron dose: 49.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)