+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of endogenous Cage-GIDAnt complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | multiprotein complex / E3 ligase / metabolic enzyme / LIGASE | |||||||||
| Biological species |  | |||||||||
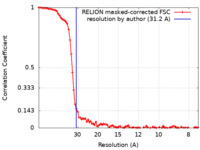
| Method | single particle reconstruction / cryo EM / Resolution: 31.2 Å | |||||||||
 Authors Authors | Sherpa D / Chrustowicz J / Qiao S / Schulman B | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Cryo-EM structures of Gid12-bound GID E3 reveal steric blockade as a mechanism inhibiting substrate ubiquitylation. Authors: Shuai Qiao / Chia-Wei Lee / Dawafuti Sherpa / Jakub Chrustowicz / Jingdong Cheng / Maximilian Duennebacke / Barbara Steigenberger / Ozge Karayel / Duc Tung Vu / Susanne von Gronau / Matthias ...Authors: Shuai Qiao / Chia-Wei Lee / Dawafuti Sherpa / Jakub Chrustowicz / Jingdong Cheng / Maximilian Duennebacke / Barbara Steigenberger / Ozge Karayel / Duc Tung Vu / Susanne von Gronau / Matthias Mann / Florian Wilfling / Brenda A Schulman /    Abstract: Protein degradation, a major eukaryotic response to cellular signals, is subject to numerous layers of regulation. In yeast, the evolutionarily conserved GID E3 ligase mediates glucose-induced ...Protein degradation, a major eukaryotic response to cellular signals, is subject to numerous layers of regulation. In yeast, the evolutionarily conserved GID E3 ligase mediates glucose-induced degradation of fructose-1,6-bisphosphatase (Fbp1), malate dehydrogenase (Mdh2), and other gluconeogenic enzymes. "GID" is a collection of E3 ligase complexes; a core scaffold, RING-type catalytic core, and a supramolecular assembly module together with interchangeable substrate receptors select targets for ubiquitylation. However, knowledge of additional cellular factors directly regulating GID-type E3s remains rudimentary. Here, we structurally and biochemically characterize Gid12 as a modulator of the GID E3 ligase complex. Our collection of cryo-EM reconstructions shows that Gid12 forms an extensive interface sealing the substrate receptor Gid4 onto the scaffold, and remodeling the degron binding site. Gid12 also sterically blocks a recruited Fbp1 or Mdh2 from the ubiquitylation active sites. Our analysis of the role of Gid12 establishes principles that may more generally underlie E3 ligase regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14338.map.gz emd_14338.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14338-v30.xml emd-14338-v30.xml emd-14338.xml emd-14338.xml | 14.7 KB 14.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14338_fsc.xml emd_14338_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_14338.png emd_14338.png | 19.8 KB | ||
| Masks |  emd_14338_msk_1.map emd_14338_msk_1.map | 58.2 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14338.cif.gz emd-14338.cif.gz | 4.1 KB | ||
| Others |  emd_14338_additional_1.map.gz emd_14338_additional_1.map.gz emd_14338_half_map_1.map.gz emd_14338_half_map_1.map.gz emd_14338_half_map_2.map.gz emd_14338_half_map_2.map.gz | 7.7 MB 45.1 MB 45.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14338 http://ftp.pdbj.org/pub/emdb/structures/EMD-14338 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14338 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14338 | HTTPS FTP |
-Validation report
| Summary document |  emd_14338_validation.pdf.gz emd_14338_validation.pdf.gz | 609.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14338_full_validation.pdf.gz emd_14338_full_validation.pdf.gz | 609 KB | Display | |
| Data in XML |  emd_14338_validation.xml.gz emd_14338_validation.xml.gz | 14.4 KB | Display | |
| Data in CIF |  emd_14338_validation.cif.gz emd_14338_validation.cif.gz | 20.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14338 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14338 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14338 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14338 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14338.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14338.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.77 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14338_msk_1.map emd_14338_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: 3D class
| File | emd_14338_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D class | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14338_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_14338_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Gid1, Gid2, Gid5, Gid7, Gid8, Gid9
| Entire | Name: Gid1, Gid2, Gid5, Gid7, Gid8, Gid9 |
|---|---|
| Components |
|
-Supramolecule #1: Gid1, Gid2, Gid5, Gid7, Gid8, Gid9
| Supramolecule | Name: Gid1, Gid2, Gid5, Gid7, Gid8, Gid9 / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 59.2 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)