[English] 日本語
 Yorodumi
Yorodumi- EMDB-14271: Structure of the SARS-CoV-2 spike glycoprotein in complex with th... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment (local refinement) | ||||||||||||
 Map data Map data | DeepEMhancer sharpened map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Coronavirus / Spike / Antibody / Complex / VIRAL PROTEIN | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
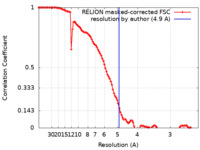
| Method | single particle reconstruction / cryo EM / Resolution: 4.9 Å | ||||||||||||
 Authors Authors | Hurdiss DL / Drulyte I | ||||||||||||
| Funding support | European Union,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Sci Immunol / Year: 2022 Journal: Sci Immunol / Year: 2022Title: An ACE2-blocking antibody confers broad neutralization and protection against Omicron and other SARS-CoV-2 variants of concern. Authors: Wenjuan Du / Daniel L Hurdiss / Dubravka Drabek / Anna Z Mykytyn / Franziska K Kaiser / Mariana González-Hernández / Diego Muñoz-Santos / Mart M Lamers / Rien van Haperen / Wentao Li / ...Authors: Wenjuan Du / Daniel L Hurdiss / Dubravka Drabek / Anna Z Mykytyn / Franziska K Kaiser / Mariana González-Hernández / Diego Muñoz-Santos / Mart M Lamers / Rien van Haperen / Wentao Li / Ieva Drulyte / Chunyan Wang / Isabel Sola / Federico Armando / Georg Beythien / Malgorzata Ciurkiewicz / Wolfgang Baumgärtner / Kate Guilfoyle / Tony Smits / Joline van der Lee / Frank J M van Kuppeveld / Geert van Amerongen / Bart L Haagmans / Luis Enjuanes / Albert D M E Osterhaus / Frank Grosveld / Berend-Jan Bosch /     Abstract: The ongoing evolution of SARS-CoV-2 has resulted in the emergence of Omicron, which displays notable immune escape potential through mutations at key antigenic sites on the spike protein. Many of ...The ongoing evolution of SARS-CoV-2 has resulted in the emergence of Omicron, which displays notable immune escape potential through mutations at key antigenic sites on the spike protein. Many of these mutations localize to the spike protein ACE2 receptor binding domain, annulling the neutralizing activity of therapeutic antibodies that were effective against other variants of concern (VOCs) earlier in the pandemic. Here, we identified a receptor-blocking human monoclonal antibody, 87G7, that retained potent in vitro neutralizing activity against SARS-CoV-2 variants including the Alpha, Beta, Gamma, Delta, and Omicron (BA.1/BA.2) VOCs. Using cryo-electron microscopy and site-directed mutagenesis experiments, we showed that 87G7 targets a patch of hydrophobic residues in the ACE2-binding site that are highly conserved in SARS-CoV-2 variants, explaining its broad neutralization capacity. 87G7 protected mice and hamsters prophylactically against challenge with all current SARS-CoV-2 VOCs and showed therapeutic activity against SARS-CoV-2 challenge in both animal models. Our findings demonstrate that 87G7 holds promise as a prophylactic or therapeutic agent for COVID-19 that is more resilient to SARS-CoV-2 antigenic diversity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14271.map.gz emd_14271.map.gz | 92.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14271-v30.xml emd-14271-v30.xml emd-14271.xml emd-14271.xml | 29.2 KB 29.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14271_fsc.xml emd_14271_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_14271.png emd_14271.png | 70.2 KB | ||
| Masks |  emd_14271_msk_1.map emd_14271_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14271.cif.gz emd-14271.cif.gz | 7.3 KB | ||
| Others |  emd_14271_additional_1.map.gz emd_14271_additional_1.map.gz emd_14271_additional_2.map.gz emd_14271_additional_2.map.gz emd_14271_additional_3.map.gz emd_14271_additional_3.map.gz emd_14271_half_map_1.map.gz emd_14271_half_map_1.map.gz emd_14271_half_map_2.map.gz emd_14271_half_map_2.map.gz | 60.9 MB 50.4 MB 97 MB 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14271 http://ftp.pdbj.org/pub/emdb/structures/EMD-14271 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14271 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14271 | HTTPS FTP |
-Validation report
| Summary document |  emd_14271_validation.pdf.gz emd_14271_validation.pdf.gz | 675.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14271_full_validation.pdf.gz emd_14271_full_validation.pdf.gz | 675.4 KB | Display | |
| Data in XML |  emd_14271_validation.xml.gz emd_14271_validation.xml.gz | 17.2 KB | Display | |
| Data in CIF |  emd_14271_validation.cif.gz emd_14271_validation.cif.gz | 22.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14271 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14271 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14271 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14271 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14271.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14271.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.24 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14271_msk_1.map emd_14271_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
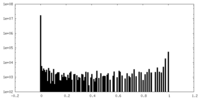
| Density Histograms |
-Additional map: Local resolution filtered map
| File | emd_14271_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution filtered map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_14271_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened map
| File | emd_14271_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_14271_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_14271_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody F...
| Entire | Name: SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody F...
| Supramolecule | Name: SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
-Supramolecule #2: SARS-CoV-2 spike protein
| Supramolecule | Name: SARS-CoV-2 spike protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Fab
| Supramolecule | Name: Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: SARS-CoV-2 spike protein
| Macromolecule | Name: SARS-CoV-2 spike protein / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF LGVYYHKNNK SWMESEFRVY SSANNCTFEY ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDGV YFASTEKSNI IRGWIFGTTL DSKTQSLLIV NNATNVVIKV CEFQFCNDPF LGVYYHKNNK SWMESEFRVY SSANNCTFEY VSQPFLMDLE GKQGNFKNLR EFVFKNIDGY FKIYSKHTPI NLVRDLPQGF SALEPLVDLP IGINITRFQT LLALHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIYQTSNFRV QPTESIVRFP NITNLCPFGE VFNATRFASV YAWNRKRISN CVADYSVLYN SASFSTFKCY GVSPTKLNDL CFTNVYADSF VIRGDEVRQI APGQTGKIAD YNYKLPDDFT GCVIAWNSNN LDSKVGGNYN YLYRLFRKSN LKPFERDIST EIYQAGSTPC NGVEGFNCYF PLQSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCGPKKS TNLVKNKCVN FNFNGLTGTG VLTESNKKFL PFQQFGRDIA DTTDAVRDPQ TLEILDITPC SFGGVSVITP GTNTSNQVAV LYQDVNCTEV PVAIHADQLT PTWRVYSTGS NVFQTRAGCL IGAEHVNNSY ECDIPIGAGI CASYQTQTNS PAAARSVASQ SIIAYTMSLG AENSVAYSNN SIAIPTNFTI SVTTEILPVS MTKTSVDCTM YICGDSTECS NLLLQYGSFC TQLNRALTGI AVEQDKNTQE VFAQVKQIYK TPPIKDFGGF NFSQILPDPS KPSKRSPIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TITSGWTFGA GPALQIPFPM QMAYRFNGIG VTQNVLYENQ KLIANQFNSA IGKIQDSLSS TPSALGKLQD VVNQNAQALN TLVKQLSSNF GAISSVLNDI LSRLDPPEAE VQIDRLITGR LQSLQTYVTQ QLIRAAEIRA SANLAATKMS ECVLGQSKRV DFCGKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVSGNCDVVI GIVNNTVYDP LQPELDSFKE ELDKYFKNHT SPDVDLGDIS GINASVVNIQ KEIDRLNEVA KNLNESLIDL QELGKYEQGS GYIPEAPRDG QAYVRKDGEW VLLSTFLGRS LEVLFQGPGH HHHHHHHSAW SHPQFEKGGG SGGGGSGGSA WSHPQFEK |
-Macromolecule #2: 87G7 heavy chain variable domain
| Macromolecule | Name: 87G7 heavy chain variable domain / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLLESGGG LVQPGGSLRL SCVASGFTFS SYVMSWVRQA PGKGLEWVAA ISGSGGNTYY ADSVKGRFTI SRDNSKNTLY LQMNSLRAED TAVYYCAKEG TYYYGSGSFW GQGTLVTVSS |
-Macromolecule #3: 87G7 light chain variable domain
| Macromolecule | Name: 87G7 light chain variable domain / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPAT LSLSPGERAT LSCRASQSFH NYLAWYQQKP GRAPRLLIFD ASNRATGIPA RFSGSGSGTD FTLTISSLEP EDFAVYYCQQ RFNWPLTFGG GTKVEIK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 2 / Number real images: 3831 / Average electron dose: 51.5 e/Å2 Details: 1331 images collected from grid 1 (0.02% FOM) and 2500 images collected from grid 2 (0% FOM) |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)