+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Fibrillin-1 microfibril arm-region | |||||||||
 Map data Map data | Fibrillin microfibril arm-region | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Microfibril / Extra-Cellular-Matrix / STRUCTURAL PROTEIN | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 18.3 Å | |||||||||
 Authors Authors | Godwin ARF / Thomson J / Holmes DF / Adamo CS / Sengle G / Sherratt MJ / Roseman AM / Baldock C | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Fibrillin microfibril structure identifies long-range effects of inherited pathogenic mutations affecting a key regulatory latent TGFβ-binding site. Authors: Alan R F Godwin / Rana Dajani / Xinyang Zhang / Jennifer Thomson / David F Holmes / Christin S Adamo / Gerhard Sengle / Michael J Sherratt / Alan M Roseman / Clair Baldock /   Abstract: Genetic mutations in fibrillin microfibrils cause serious inherited diseases, such as Marfan syndrome and Weill-Marchesani syndrome (WMS). These diseases typically show major dysregulation of tissue ...Genetic mutations in fibrillin microfibrils cause serious inherited diseases, such as Marfan syndrome and Weill-Marchesani syndrome (WMS). These diseases typically show major dysregulation of tissue development and growth, particularly in skeletal long bones, but links between the mutations and the diseases are unknown. Here we describe a detailed structural analysis of native fibrillin microfibrils from mammalian tissue by cryogenic electron microscopy. The major bead region showed pseudo eightfold symmetry where the amino and carboxy termini reside. On the basis of this structure, we show that a WMS deletion mutation leads to the induction of a structural rearrangement that blocks interaction with latent TGFβ-binding protein-1 at a remote site. Separate deletion of this binding site resulted in the assembly of shorter fibrillin microfibrils with structural alterations. The integrin αβ-binding site was also mapped onto the microfibril structure. These results establish that in complex extracellular assemblies, such as fibrillin microfibrils, mutations may have long-range structural consequences leading to the disruption of growth factor signaling and the development of disease. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13986.map.gz emd_13986.map.gz | 48 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13986-v30.xml emd-13986-v30.xml emd-13986.xml emd-13986.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13986_fsc.xml emd_13986_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_13986.png emd_13986.png | 23.2 KB | ||
| Filedesc metadata |  emd-13986.cif.gz emd-13986.cif.gz | 4.6 KB | ||
| Others |  emd_13986_additional_1.map.gz emd_13986_additional_1.map.gz | 47.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13986 http://ftp.pdbj.org/pub/emdb/structures/EMD-13986 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13986 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13986 | HTTPS FTP |
-Validation report
| Summary document |  emd_13986_validation.pdf.gz emd_13986_validation.pdf.gz | 458.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_13986_full_validation.pdf.gz emd_13986_full_validation.pdf.gz | 457.6 KB | Display | |
| Data in XML |  emd_13986_validation.xml.gz emd_13986_validation.xml.gz | 11 KB | Display | |
| Data in CIF |  emd_13986_validation.cif.gz emd_13986_validation.cif.gz | 14.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13986 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13986 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13986 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13986 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13986.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13986.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Fibrillin microfibril arm-region | ||||||||||||||||||||
| Voxel size | X=Y=Z: 2.22 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Fibrillin microfibril arm-region with bread region density removed
| File | emd_13986_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Fibrillin microfibril arm-region with bread region density removed | ||||||||||||
| Projections & Slices |
| ||||||||||||
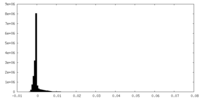
| Density Histograms |
- Sample components
Sample components
-Entire : Fibrillin-1 microfibril arm region
| Entire | Name: Fibrillin-1 microfibril arm region |
|---|---|
| Components |
|
-Supramolecule #1: Fibrillin-1 microfibril arm region
| Supramolecule | Name: Fibrillin-1 microfibril arm region / type: tissue / ID: 1 / Parent: 0 Details: Microfibrils generated from sonication of bovine ciliary zonule tissue. |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Solutions were made fresh and filtered using 0.2 um filters. | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Blot for 4 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4000 pixel / Digitization - Dimensions - Height: 4000 pixel / Digitization - Frames/image: 1-20 / Number grids imaged: 1 / Average exposure time: 20.0 sec. / Average electron dose: 66.0 e/Å2 / Details: Movies were collected in counting mode |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X