+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the NLRP3 PYD filament | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NLRP3 / Pyrin domain / filament / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationdetection of biotic stimulus / molecular sensor activity / phosphatidylinositol phosphate binding / positive regulation of T-helper 2 cell differentiation / positive regulation of T-helper 2 cell cytokine production / interphase microtubule organizing center / positive regulation of type 2 immune response / NLRP3 inflammasome complex / NLRP3 inflammasome complex assembly / peptidoglycan binding ...detection of biotic stimulus / molecular sensor activity / phosphatidylinositol phosphate binding / positive regulation of T-helper 2 cell differentiation / positive regulation of T-helper 2 cell cytokine production / interphase microtubule organizing center / positive regulation of type 2 immune response / NLRP3 inflammasome complex / NLRP3 inflammasome complex assembly / peptidoglycan binding / cysteine-type endopeptidase activator activity / phosphatidylinositol-4-phosphate binding / osmosensory signaling pathway / negative regulation of non-canonical NF-kappaB signal transduction / negative regulation of interleukin-1 beta production / pattern recognition receptor signaling pathway / positive regulation of interleukin-4 production / negative regulation of acute inflammatory response / microtubule organizing center / The NLRP3 inflammasome / pyroptotic inflammatory response / Purinergic signaling in leishmaniasis infection / signaling adaptor activity / : / positive regulation of interleukin-1 beta production / molecular condensate scaffold activity / defense response / Cytoprotection by HMOX1 / protein maturation / ADP binding / cellular response to virus / protein homooligomerization / positive regulation of non-canonical NF-kappaB signal transduction / negative regulation of inflammatory response / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / Metalloprotease DUBs / positive regulation of inflammatory response / SARS-CoV-1 activates/modulates innate immune responses / cellular response to lipopolysaccharide / regulation of inflammatory response / DNA-binding transcription factor binding / sequence-specific DNA binding / molecular adaptor activity / protein-macromolecule adaptor activity / inflammatory response / Golgi membrane / innate immune response / apoptotic process / SARS-CoV-2 activates/modulates innate and adaptive immune responses / endoplasmic reticulum / signal transduction / ATP hydrolysis activity / positive regulation of transcription by RNA polymerase II / mitochondrion / extracellular region / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Hochheiser IV / Hagelueken G | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Directionality of PYD filament growth determined by the transition of NLRP3 nucleation seeds to ASC elongation. Authors: Inga V Hochheiser / Heide Behrmann / Gregor Hagelueken / Juan F Rodríguez-Alcázar / Anja Kopp / Eicke Latz / Elmar Behrmann / Matthias Geyer /   Abstract: Inflammasomes sense intrinsic and extrinsic danger signals to trigger inflammatory responses and pyroptotic cell death. Homotypic pyrin domain (PYD) interactions of inflammasome forming nucleotide- ...Inflammasomes sense intrinsic and extrinsic danger signals to trigger inflammatory responses and pyroptotic cell death. Homotypic pyrin domain (PYD) interactions of inflammasome forming nucleotide-binding oligomerization domain (NOD)-like receptors with the adaptor protein ASC (apoptosis-associated speck-like protein containing a CARD) mediate oligomerization into filamentous assemblies. We describe the cryo-electron microscopy (cryo-EM) structure of the human NLRP3 filament and identify a pattern of highly polar interface residues that form the homomeric interactions leading to characteristic filament ends designated as A- and B-ends. Coupling a titration polymerization assay to cryo-EM, we demonstrate that ASC adaptor protein elongation on NLRP3 nucleation seeds is unidirectional, associating exclusively to the B-end of the filament. Notably, NLRP3 and ASC PYD filaments exhibit the same symmetry in rotation and axial rise per subunit, allowing a continuous transition between NLRP3 and ASC. Integrating the directionality of filament growth, we present a molecular model of the ASC speck consisting of active NLRP3, ASC, and Caspase-1 proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13727.map.gz emd_13727.map.gz | 26.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13727-v30.xml emd-13727-v30.xml emd-13727.xml emd-13727.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13727.png emd_13727.png | 77.3 KB | ||
| Filedesc metadata |  emd-13727.cif.gz emd-13727.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13727 http://ftp.pdbj.org/pub/emdb/structures/EMD-13727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13727 | HTTPS FTP |
-Related structure data
| Related structure data |  7pzdMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13727.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13727.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.067 Å | ||||||||||||||||||||||||||||||||||||
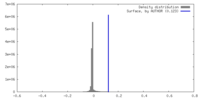
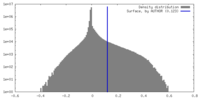
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : filament formed by the human NLRP3 Pyrin domain
| Entire | Name: filament formed by the human NLRP3 Pyrin domain |
|---|---|
| Components |
|
-Supramolecule #1: filament formed by the human NLRP3 Pyrin domain
| Supramolecule | Name: filament formed by the human NLRP3 Pyrin domain / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: NACHT, LRR and PYD domains-containing protein 3
| Macromolecule | Name: NACHT, LRR and PYD domains-containing protein 3 / type: protein_or_peptide / ID: 1 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.486243 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASTRCKLAR YLEDLEDVDL KKFKMHLEDY PPQKGCIPLP RGQTEKADHV DLATLMIDFN GEEKAWAMAV WIFAAINRRD LYEKAKRDE PKWGSDNARV SNPTVISQE UniProtKB: NACHT, LRR and PYD domains-containing protein 3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20 mM HEPES (pH 7.5), 150 mM NaCl, 0.5 mM TCEP |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 293 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 14.32 Å Applied symmetry - Helical parameters - Δ&Phi: 54.88 ° Applied symmetry - Helical parameters - Axial symmetry: C3 (3 fold cyclic) Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 60333 |
|---|---|
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)