[English] 日本語
 Yorodumi
Yorodumi- EMDB-13586: Cryo-electron tomography of ASC signalling sites in pyroptotic ce... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron tomography of ASC signalling sites in pyroptotic cells (2) | |||||||||
 Map data Map data | Cryo-ET of ASC-mCerulean speck in iBMDMs | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ASC-mCerulean speck / PROTEIN FIBRIL | |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Liu Y / Modis Y / Alemayehu H / Hopkins LJ / Borgeaud AC / Heroven C / Howe JD / Boulanger J / Bryant CE / Zhai H | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-electron tomography of NLRP3-activated ASC complexes reveals organelle co-localization. Authors: Yangci Liu / Haoming Zhai / Helen Alemayehu / Jérôme Boulanger / Lee J Hopkins / Alicia C Borgeaud / Christina Heroven / Jonathan D Howe / Kendra E Leigh / Clare E Bryant / Yorgo Modis /   Abstract: NLRP3 induces caspase-1-dependent pyroptotic cell death to drive inflammation. Aberrant activity of NLRP3 occurs in many human diseases. NLRP3 activation induces ASC polymerization into a single, ...NLRP3 induces caspase-1-dependent pyroptotic cell death to drive inflammation. Aberrant activity of NLRP3 occurs in many human diseases. NLRP3 activation induces ASC polymerization into a single, micron-scale perinuclear punctum. Higher resolution imaging of this signaling platform is needed to understand how it induces pyroptosis. Here, we apply correlative cryo-light microscopy and cryo-electron tomography to visualize ASC/caspase-1 in NLRP3-activated cells. The puncta are composed of branched ASC filaments, with a tubular core formed by the pyrin domain. Ribosomes and Golgi-like or endosomal vesicles permeate the filament network, consistent with roles for these organelles in NLRP3 activation. Mitochondria are not associated with ASC but have outer-membrane discontinuities the same size as gasdermin D pores, consistent with our data showing gasdermin D associates with mitochondria and contributes to mitochondrial depolarization. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13586.map.gz emd_13586.map.gz | 416 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13586-v30.xml emd-13586-v30.xml emd-13586.xml emd-13586.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13586.png emd_13586.png | 237.8 KB | ||
| Filedesc metadata |  emd-13586.cif.gz emd-13586.cif.gz | 4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13586 http://ftp.pdbj.org/pub/emdb/structures/EMD-13586 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13586 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13586 | HTTPS FTP |
-Related structure data
| Related structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13586.map.gz / Format: CCP4 / Size: 450 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13586.map.gz / Format: CCP4 / Size: 450 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
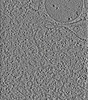
| Annotation | Cryo-ET of ASC-mCerulean speck in iBMDMs | ||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 13.58 Å | ||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : FIB milled iBMDMs
| Entire | Name: FIB milled iBMDMs |
|---|---|
| Components |
|
-Supramolecule #1: FIB milled iBMDMs
| Supramolecule | Name: FIB milled iBMDMs / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Details: DMEM medium supplemented with 10% heat-inactivated FBS (Gibco) |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: LEICA EM GP |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 1 / Focused ion beam - Duration: 1500 / Focused ion beam - Temperature: 83 K / Focused ion beam - Initial thickness: 10000 / Focused ion beam - Final thickness: 200 Focused ion beam - Details: Lamella were made using a Scios DualBeam FIB/SEM (FEI) equipped with a Quorum cryo-stage (PP3010T). Rough milling was performed using 0.5 to 0.1 nA current. Fine milling ...Focused ion beam - Details: Lamella were made using a Scios DualBeam FIB/SEM (FEI) equipped with a Quorum cryo-stage (PP3010T). Rough milling was performed using 0.5 to 0.1 nA current. Fine milling was done using 50 to 10 pA current.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 121 / Average electron dose: 1.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 7.0 µm / Nominal defocus min: 4.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution method: OTHER / Software - Name: eTomo / Number images used: 99 |
|---|
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
















