[English] 日本語
 Yorodumi
Yorodumi- EMDB-1192: Conformational changes in the AAA ATPase p97-p47 adaptor complex. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1192 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
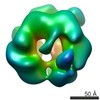
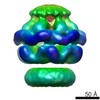
| Title | Conformational changes in the AAA ATPase p97-p47 adaptor complex. | |||||||||
 Map data Map data | AAA ATPase p97 in complex with p47 adaptor, incubated with ADP | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 27.0 Å | |||||||||
 Authors Authors | Beuron F / Dreveny I / Yuan X / Pye VE / McKeown C / Briggs LC / Cliff MJ / Kaneko Y / Wallis R / Isaacson RL ...Beuron F / Dreveny I / Yuan X / Pye VE / McKeown C / Briggs LC / Cliff MJ / Kaneko Y / Wallis R / Isaacson RL / Ladbury JE / Matthews SJ / Kondo H / Zhang X / Freemont PS | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2006 Journal: EMBO J / Year: 2006Title: Conformational changes in the AAA ATPase p97-p47 adaptor complex. Authors: Fabienne Beuron / Ingrid Dreveny / Xuemei Yuan / Valerie E Pye / Ciaran McKeown / Louise C Briggs / Matthew J Cliff / Yayoi Kaneko / Russell Wallis / Rivka L Isaacson / John E Ladbury / ...Authors: Fabienne Beuron / Ingrid Dreveny / Xuemei Yuan / Valerie E Pye / Ciaran McKeown / Louise C Briggs / Matthew J Cliff / Yayoi Kaneko / Russell Wallis / Rivka L Isaacson / John E Ladbury / Steve J Matthews / Hisao Kondo / Xiaodong Zhang / Paul S Freemont /  Abstract: The AAA+ATPase p97/VCP, helped by adaptor proteins, exerts its essential role in cellular events such as endoplasmic reticulum-associated protein degradation or the reassembly of Golgi, ER and the ...The AAA+ATPase p97/VCP, helped by adaptor proteins, exerts its essential role in cellular events such as endoplasmic reticulum-associated protein degradation or the reassembly of Golgi, ER and the nuclear envelope after mitosis. Here, we report the three-dimensional cryo-electron microscopy structures at approximately 20 Angstroms resolution in two nucleotide states of the endogenous hexameric p97 in complex with a recombinant p47 trimer, one of the major p97 adaptor proteins involved in membrane fusion. Depending on the nucleotide state, we observe the p47 trimer to be in two distinct arrangements on top of the p97 hexamer. By combining the EM data with NMR and other biophysical measurements, we propose a model of ATP-dependent p97(N) domain motions that lead to a rearrangement of p47 domains, which could result in the disassembly of target protein complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1192.map.gz emd_1192.map.gz | 12.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1192-v30.xml emd-1192-v30.xml emd-1192.xml emd-1192.xml | 10.5 KB 10.5 KB | Display Display |  EMDB header EMDB header |
| Images |  1192.gif 1192.gif | 28.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1192 http://ftp.pdbj.org/pub/emdb/structures/EMD-1192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1192 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1192 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1192.map.gz / Format: CCP4 / Size: 12.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1192.map.gz / Format: CCP4 / Size: 12.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AAA ATPase p97 in complex with p47 adaptor, incubated with ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Rat endogenous AAA ATPase p97 bound to p47, 'ADP' form
| Entire | Name: Rat endogenous AAA ATPase p97 bound to p47, 'ADP' form |
|---|---|
| Components |
|
-Supramolecule #1000: Rat endogenous AAA ATPase p97 bound to p47, 'ADP' form
| Supramolecule | Name: Rat endogenous AAA ATPase p97 bound to p47, 'ADP' form type: sample / ID: 1000 Oligomeric state: one homohexamer of p97 bound to one trimer of p47 Number unique components: 2 |
|---|---|
| Molecular weight | Experimental: 650 KDa / Theoretical: 720 KDa / Method: STEM |
-Macromolecule #1: p97
| Macromolecule | Name: p97 / type: protein_or_peptide / ID: 1 / Name.synonym: VCP / Number of copies: 6 / Oligomeric state: homohexamer / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 97 MDa / Theoretical: 89 MDa |
-Macromolecule #2: p47
| Macromolecule | Name: p47 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Oligomeric state: trimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 47 MDa / Theoretical: 41 MDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM Tris-HCl, 150 mM KCl, 1 mM DTT, pH 7.4 |
| Grid | Details: lacey carbon film |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA KF80 / Details: Vitrification instrument: Reichert KF 80 plunger / Method: blot for 2 seconds befre plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Date | Jun 6, 2002 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Digitization - Sampling interval: 6.35 µm / Number real images: 4 / Average electron dose: 15 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.4 mm / Nominal defocus max: 1.9 µm / Nominal defocus min: 1.1 µm / Nominal magnification: 38000 |
| Sample stage | Specimen holder: eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 27.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Imagic / Number images used: 2700 |
| Final two d classification | Number classes: 44 |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















