[English] 日本語
 Yorodumi
Yorodumi- EMDB-1155: The mechanism of HIV-1 core assembly: insights from three-dimensi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1155 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The mechanism of HIV-1 core assembly: insights from three-dimensional reconstructions of authentic virions. | |||||||||
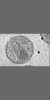
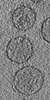
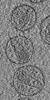
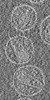
 Map data Map data | Tomographic reconstruction of HIV-1 virions | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human immunodeficiency virus Human immunodeficiency virus | |||||||||
| Method | electron tomography / cryo EM / negative staining | |||||||||
 Authors Authors | Briggs JAG / Grunewald K / Glass B / Forster F | |||||||||
 Citation Citation |  Journal: Structure / Year: 2006 Journal: Structure / Year: 2006Title: The mechanism of HIV-1 core assembly: insights from three-dimensional reconstructions of authentic virions. Authors: John A G Briggs / Kay Grünewald / Bärbel Glass / Friedrich Förster / Hans-Georg Kräusslich / Stephen D Fuller /  Abstract: Infectious HIV particles contain a characteristic cone-shaped core encasing the viral RNA and replication proteins. The core exhibits significant heterogeneity in size and shape, yet consistently ...Infectious HIV particles contain a characteristic cone-shaped core encasing the viral RNA and replication proteins. The core exhibits significant heterogeneity in size and shape, yet consistently forms a well-defined structure. The mechanism by which the core is assembled in the maturing virion remains poorly understood. Using cryo-electron tomography, we have produced three-dimensional reconstructions of authentic, unstained HIV-1. These reveal the viral morphology with unprecedented clarity and suggest the following mechanism for core formation inside the extracellular virion: core growth initiates at the narrow end of the cone and proceeds toward the distal side of the virion until limited by the viral membrane. Curvature and closure of the broad end of the core are then directed by the inner surface of the viral membrane. This mechanism accommodates significant flexibility in lattice growth while ensuring the closure of cores of variable size and shape. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1155.map.gz emd_1155.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1155-v30.xml emd-1155-v30.xml emd-1155.xml emd-1155.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  1155.gif 1155.gif | 33 KB | ||
| Others |  emd_1155_additional_1.map.gz emd_1155_additional_1.map.gz | 3.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1155 http://ftp.pdbj.org/pub/emdb/structures/EMD-1155 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1155 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1155 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1155.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS SIGNED BYTE Download / File: emd_1155.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS SIGNED BYTE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tomographic reconstruction of HIV-1 virions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 16.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 1155 additional 1.map
| File | emd_1155_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HIV-1 virions
| Entire | Name: HIV-1 virions |
|---|---|
| Components |
|
-Supramolecule #1000: HIV-1 virions
| Supramolecule | Name: HIV-1 virions / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Human immunodeficiency virus
| Supramolecule | Name: Human immunodeficiency virus / type: virus / ID: 1 / Name.synonym: HIV-1 / NCBI-ID: 12721 / Sci species name: Human immunodeficiency virus / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: HIV-1 |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | electron tomography |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 / Details: PBS |
|---|---|
| Staining | Type: NEGATIVE / Details: unstained |
| Grid | Details: holey carbon film |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Specialist optics | Energy filter - Name: GIF2000 / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Oct 22, 2004 |
| Image recording | Average electron dose: 50 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 36500 / Illumination mode: FLOOD BEAM / Imaging mode: OTHER / Cs: 2.0 mm / Nominal defocus max: 6.5 µm / Nominal defocus min: 5.5 µm |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: 68 ° / Tilt series - Axis1 - Max angle: 68 ° / Tilt series - Axis1 - Angle increment: 2 ° |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Software - Name:  IMOD IMOD |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)