[English] 日本語
 Yorodumi
Yorodumi- EMDB-10894: PorLM complex from Porphyromonas gingivalis solubilised in LMNG m... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10894 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | PorLM complex from Porphyromonas gingivalis solubilised in LMNG micelle | ||||||||||||
 Map data Map data | Volume of full length PorLM in LMNG micelle | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |  Flavobacterium johnsoniae (bacteria) / Flavobacterium johnsoniae (bacteria) /  Porphyromonas gingivalis (bacteria) / Porphyromonas gingivalis (bacteria) /  Porphyromonas gingivalis (strain ATCC 33277 / DSM 20709 / CIP 103683 / JCM 12257 / NCTC 11834 / 2561) (bacteria) Porphyromonas gingivalis (strain ATCC 33277 / DSM 20709 / CIP 103683 / JCM 12257 / NCTC 11834 / 2561) (bacteria) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.6 Å | ||||||||||||
 Authors Authors | Hennell James R / Deme JC / Lea SM | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2021 Journal: Nat Microbiol / Year: 2021Title: Structure and mechanism of the proton-driven motor that powers type 9 secretion and gliding motility. Authors: Rory Hennell James / Justin C Deme / Andreas Kjӕr / Felicity Alcock / Augustinas Silale / Frédéric Lauber / Steven Johnson / Ben C Berks / Susan M Lea /  Abstract: Three classes of ion-driven protein motors have been identified to date: ATP synthase, the bacterial flagellar motor and a proton-driven motor that powers gliding motility and the type 9 protein ...Three classes of ion-driven protein motors have been identified to date: ATP synthase, the bacterial flagellar motor and a proton-driven motor that powers gliding motility and the type 9 protein secretion system in Bacteroidetes bacteria. Here, we present cryo-electron microscopy structures of the gliding motility/type 9 protein secretion system motors GldLM from Flavobacterium johnsoniae and PorLM from Porphyromonas gingivalis. The motor is an asymmetric inner membrane protein complex in which the single transmembrane helices of two periplasm-spanning GldM/PorM proteins are positioned inside a ring of five GldL/PorL proteins. Mutagenesis and single-molecule tracking identify protonatable amino acid residues in the transmembrane domain of the complex that are important for motor function. Our data provide evidence for a mechanism in which proton flow results in rotation of the periplasm-spanning GldM/PorM dimer inside the intra-membrane GldL/PorL ring to drive processes at the bacterial outer membrane. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10894.map.gz emd_10894.map.gz | 332.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10894-v30.xml emd-10894-v30.xml emd-10894.xml emd-10894.xml | 17.6 KB 17.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10894.png emd_10894.png | 32.4 KB | ||
| Masks |  emd_10894_msk_1.map emd_10894_msk_1.map | 421.9 MB |  Mask map Mask map | |
| Others |  emd_10894_half_map_1.map.gz emd_10894_half_map_1.map.gz emd_10894_half_map_2.map.gz emd_10894_half_map_2.map.gz | 338.5 MB 338.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10894 http://ftp.pdbj.org/pub/emdb/structures/EMD-10894 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10894 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10894 | HTTPS FTP |
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10894.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10894.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Volume of full length PorLM in LMNG micelle | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
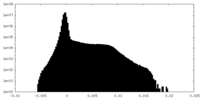
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10894_msk_1.map emd_10894_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
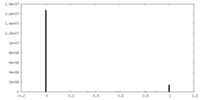
| Density Histograms |
-Half map: half map 1
| File | emd_10894_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
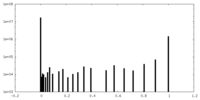
| Density Histograms |
-Half map: half map 2
| File | emd_10894_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
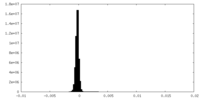
| Density Histograms |
- Sample components
Sample components
-Entire : PorLM
| Entire | Name: PorLM |
|---|---|
| Components |
|
-Supramolecule #1: PorLM
| Supramolecule | Name: PorLM / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Flavobacterium johnsoniae (bacteria) Flavobacterium johnsoniae (bacteria) |
| Recombinant expression | Organism:  |
-Macromolecule #1: PorL
| Macromolecule | Name: PorL / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Porphyromonas gingivalis (bacteria) Porphyromonas gingivalis (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MGHYRRYKNI LEMYLASHKG RRLLNIVYSW GAAVVILGAL FKLLHLPMGN EMLFVGMITE FLVFFISGF EKPAMEYHWE EVFPELDSKN PMDRREMEQR REYLREKAKE AAAYAERSSS V RLASASLG TQPQEQSKPA TPFQSQLTGI LPEEQIQRLS EGIDKLAEAG ...String: MGHYRRYKNI LEMYLASHKG RRLLNIVYSW GAAVVILGAL FKLLHLPMGN EMLFVGMITE FLVFFISGF EKPAMEYHWE EVFPELDSKN PMDRREMEQR REYLREKAKE AAAYAERSSS V RLASASLG TQPQEQSKPA TPFQSQLTGI LPEEQIQRLS EGIDKLAEAG EQLARIGRTA AA MTESYEQ MQADQEGLRL NSQSYIQQME SLSRNISGLN TIYEIQLKGI SSQIDTIDRI NRG LAHIRD MYDNSVIDSS SFRNENERMA RQLTQLNEVY ARLLQALTTN VGLPGMPGNF GASN PSSSG SSPL |
-Macromolecule #2: PorM
| Macromolecule | Name: PorM / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Porphyromonas gingivalis (strain ATCC 33277 / DSM 20709 / CIP 103683 / JCM 12257 / NCTC 11834 / 2561) (bacteria) Porphyromonas gingivalis (strain ATCC 33277 / DSM 20709 / CIP 103683 / JCM 12257 / NCTC 11834 / 2561) (bacteria) |
| Sequence | String: MAVGSNGNAN RQKMINLMYL VFIAMMALNV SSEVLDGFDK VDKSLTSSID GSDKRNNLVL SELNTAYRT NPEKVKVWYE RSLVLQKEAD SLCTFIDDLK LAIARESDGK DAKVNDIRRK D NLDASSVV MLNPINGKGS TLRKEVDKFR ELVATLMTDK AKLKLIEQAL ...String: MAVGSNGNAN RQKMINLMYL VFIAMMALNV SSEVLDGFDK VDKSLTSSID GSDKRNNLVL SELNTAYRT NPEKVKVWYE RSLVLQKEAD SLCTFIDDLK LAIARESDGK DAKVNDIRRK D NLDASSVV MLNPINGKGS TLRKEVDKFR ELVATLMTDK AKLKLIEQAL NTESGTKGKS WE SSLFENM PTVAAITLLT KLQSDVRYAQ GEVLADLVKS VDVGDYRVNS ITAQVIPQSQ IVM SGDTYK ANIVLSSVDT TQRPDVFVNG KLLSPENMGL FTATAGAPGT YPVKGYIEMM GNDG VKIRR DFESEYFVTE PMASVAPTMM NVLYAGIDNP INIAVPGVAQ QNVSATINNG TLTRR GNLW IARPTKVGSE AIISVTAQSG GRTIQMAKTT LRVRALPDPL PYIEYKDVQG NTKRFK GGR LGKREILAAG GIKAALDDDL LEVNYTVVKF QLVFYDSMGN SIPEVSDGAS FSERQKR QI QNLGKGKRFY VTEVIARGPD GIERKIPAIE VIVN |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.0003 µm / Nominal defocus min: 0.0001 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)