+ Open data
Open data
- Basic information
Basic information
| Entry | 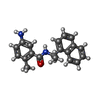 Database: PDB chemical components / ID: TTT Database: PDB chemical components / ID: TTT |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: TTT / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3E9S | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB / BindingDB /  Brenda Brenda       / /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  PubChem / PubChem /  SureChEMBL SureChEMBL   / /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 6 items

PDB-3e9s: 
A new class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication

PDB-7cjm: 
SARS CoV-2 PLpro in complex with GRL0617

PDB-7cmd: 
Crystal structure of the SARS-CoV-2 PLpro with GRL0617

PDB-7jir: 
The crystal structure of Papain-Like Protease of SARS CoV-2 , C111S mutant, in complex with PLP_Snyder457 inhibitor

PDB-7jrn: 
Crystal structure of the wild type SARS-CoV-2 papain-like protease (PLPro) with inhibitor GRL0617

PDB-7skq: 
BtSCoV-Rf1.2004 Papain-Like protease bound to the non-covalent inhibitor GRL-0617
 Movie
Movie Controller
Controller



