[English] 日本語
 Yorodumi
Yorodumi- ChemComp-SLX: (13aS)-3,10-dimethoxy-5,8,13,13a-tetrahydro-6H-isoquino[3,2-a]iso... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 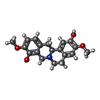 Database: PDB chemical components / ID: SLX Database: PDB chemical components / ID: SLX | ||
|---|---|---|---|
| Name | Name: ( Synonyms: (S)-scoulerine Comment | alkaloid*YM | |
-Chemical information
| Composition |  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: SLX / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3FW9 | ||||||||
| History |
| ||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB / BindingDB /  Brenda Brenda  / /  ChEBI / ChEBI /  ChemicalBook / ChemicalBook /  HMDB / HMDB /  KEGG_Ligand / KEGG_Ligand /  Metabolights / Metabolights /  Nikkaji / Nikkaji /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  Rhea / Rhea /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | (| OpenEye OEToolkits 1.5.0 | ( | |
|---|
-PDB entries
Showing all 6 items

PDB-3fw9: 
Structure of berberine bridge enzyme in complex with (S)-scoulerine

PDB-6i6k: 
Papaver somniferum O-methyltransferase 1

PDB-6i6m: 
Papaver somniferum O-methyltransferase 1

PDB-6i6n: 
Papaver somniferum O-methyltransferase 1

PDB-6neh: 
N191D, F205S mutant of scoulerine 9-O-methyltransferase from Thalictrum flavum complexed with (13aS)-3,10-dimethoxy-5,8,13,13a-tetrahydro-6H-isoquino[3,2-a]isoquinoline-2,9-diol and S-ADENOSYL-L-HOMOCYSTEINE

PDB-6nej: 
scoulerine 9-O-methyltransferase from Thalictrum flavum complexed (13aS)-3,10-dimethoxy-5,8,13,13a-tetrahydro-6H-isoquino[3,2-a]isoquinoline-2,9-diol and with S-ADENOSYL-L-HOMOCYSTEINE
 Movie
Movie Controller
Controller


