+ Open data
Open data
- Basic information
Basic information
| Entry | 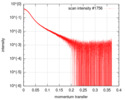 |
|---|---|
 Sample Sample | Apolipoprotein E4 (methylated) bound to 4 µM Suramin
|
| Function / homology |  Function and homology information Function and homology informationlipid transport involved in lipid storage / maintenance of location in cell / intermediate-density lipoprotein particle clearance / positive regulation of lipid transport across blood-brain barrier / regulation of cellular response to very-low-density lipoprotein particle stimulus / metal chelating activity / triglyceride-rich lipoprotein particle clearance / negative regulation of triglyceride metabolic process / discoidal high-density lipoprotein particle / Transcriptional regulation by the AP-2 (TFAP2) family of transcription factors ...lipid transport involved in lipid storage / maintenance of location in cell / intermediate-density lipoprotein particle clearance / positive regulation of lipid transport across blood-brain barrier / regulation of cellular response to very-low-density lipoprotein particle stimulus / metal chelating activity / triglyceride-rich lipoprotein particle clearance / negative regulation of triglyceride metabolic process / discoidal high-density lipoprotein particle / Transcriptional regulation by the AP-2 (TFAP2) family of transcription factors / chylomicron remnant clearance / negative regulation of cholesterol biosynthetic process / chylomicron remnant / lipoprotein particle / regulation of amyloid-beta clearance / intermediate-density lipoprotein particle / NMDA glutamate receptor clustering / very-low-density lipoprotein particle remodeling / Chylomicron clearance / acylglycerol homeostasis / phosphatidylcholine-sterol O-acyltransferase activator activity / positive regulation of phospholipid efflux / lipid carrier activity / positive regulation of low-density lipoprotein particle receptor catabolic process / Chylomicron remodeling / cellular response to lipoprotein particle stimulus / very-low-density lipoprotein particle clearance / Chylomicron assembly / high-density lipoprotein particle clearance / phospholipid efflux / lipoprotein catabolic process / very-low-density lipoprotein particle receptor binding / regulation of protein metabolic process / chylomicron / high-density lipoprotein particle remodeling / multivesicular body, internal vesicle / positive regulation of amyloid-beta clearance / reverse cholesterol transport / positive regulation of cholesterol metabolic process / regulation of amyloid fibril formation / high-density lipoprotein particle assembly / host-mediated activation of viral process / lipoprotein biosynthetic process / melanosome organization / cholesterol transfer activity / regulation of behavioral fear response / low-density lipoprotein particle / cholesterol catabolic process / protein import / high-density lipoprotein particle / very-low-density lipoprotein particle / low-density lipoprotein particle remodeling / amyloid precursor protein metabolic process / response to caloric restriction / heparan sulfate proteoglycan binding / negative regulation of amyloid fibril formation / synaptic transmission, cholinergic / positive regulation of membrane protein ectodomain proteolysis / regulation of amyloid precursor protein catabolic process / regulation of Cdc42 protein signal transduction / negative regulation of endothelial cell migration / HDL remodeling / artery morphogenesis / negative regulation of protein metabolic process / regulation of cholesterol metabolic process / cholesterol efflux / regulation of axon extension / Scavenging by Class A Receptors / triglyceride metabolic process / triglyceride homeostasis / low-density lipoprotein particle receptor binding / positive regulation of amyloid fibril formation / virion assembly / regulation of innate immune response / negative regulation of endothelial cell proliferation / negative regulation of platelet-derived growth factor receptor signaling pathway / antioxidant activity / positive regulation of lipoprotein transport / response to dietary excess / positive regulation of dendritic spine development / negative regulation of amyloid-beta formation / lipoprotein particle binding / AMPA glutamate receptor clustering / negative regulation of blood vessel endothelial cell migration / negative regulation of long-term synaptic potentiation / negative regulation of platelet activation / locomotory exploration behavior / negative regulation of blood coagulation / positive regulation of dendritic spine maintenance / negative regulation of protein secretion / positive regulation of cholesterol efflux / fatty acid homeostasis / intracellular transport / regulation of neuronal synaptic plasticity / regulation of protein-containing complex assembly / positive regulation of endocytosis / long-chain fatty acid transport / Nuclear signaling by ERBB4 / positive regulation of lipid biosynthetic process / long-term memory Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDGV8 SASDGV8 |
|---|
-Related structure data
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
|---|
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: Apolipoprotein E4 (methylated) bound to 4 µM Suramin / Specimen concentration: 4 mg/ml / Entity id: 1109 / 1682 |
|---|---|
| Buffer | Name: 20 mM HEPES, 300 mM NaCl, 1 mM TCEP / pH: 8 |
| Entity #1109 | Name: ApoE4 / Type: protein / Description: Apolipoprotein E4 / Formula weight: 34.712 / Num. of mol.: 4 / Source: Homo sapiens / References: UniProt: P02649 Sequence: GPHMKVEQAV ETEPEPELRQ QTEWQSGQRW ELALGRFWDY LRWVQTLSEQ VQEELLSSQV TQELRALMDE TMKELKAYKS ELEEQLTPVA EETRARLSKE LQAAQARLGA DMEDVRGRLV QYRGEVQAML GQSTEELRVR LASHLRKLRK RLLRDADDLQ KRLAVYQAGA ...Sequence: GPHMKVEQAV ETEPEPELRQ QTEWQSGQRW ELALGRFWDY LRWVQTLSEQ VQEELLSSQV TQELRALMDE TMKELKAYKS ELEEQLTPVA EETRARLSKE LQAAQARLGA DMEDVRGRLV QYRGEVQAML GQSTEELRVR LASHLRKLRK RLLRDADDLQ KRLAVYQAGA REGAERGLSA IRERLGPLVE QGRVRAATVG SLAGQPLQER AQAWGERLRA RMEEMGSRTR DRLDEVKEQV AEVRAKLEEQ AQQIRLQAEA FQARLKSWFE PLVEDMQRQW AGLVEKVQAA VGTSAAPVPS DNH |
| Entity #1682 | Type: other / Description: Suramin / Formula weight: 1.297 / Num. of mol.: 1 Sequence: C51H40N6O2 3S6 |
-Experimental information
| Beam | Instrument name: Diamond Light Source B21 / City: Didcot / 国: UK  / Shape: 1 x 5 mm / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 4.036 mm / Shape: 1 x 5 mm / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 4.036 mm | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | |||||||||||||||||||||
| Scan | Measurement date: May 3, 2018 / Storage temperature: 20 °C / Cell temperature: 20 °C / Exposure time: 1 sec. / Number of frames: 28 / Unit: 1/A /
| |||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller















