[English] 日本語
 Yorodumi
Yorodumi- SASDGC3: DNA-binding protein HU-alpha, E34K mutant bound to 80 bp DNA (pH 5.5) -
+ Open data
Open data
- Basic information
Basic information
| Entry | 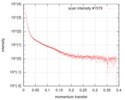 |
|---|---|
 Sample Sample | DNA-binding protein HU-alpha, E34K mutant bound to 80 bp DNA (pH 5.5)
|
| Function / homology |  Function and homology information Function and homology informationHU-DNA complex / DnaA-HU complex / bacterial nucleoid DNA packaging / chromosome condensation / DNA replication initiation / structural constituent of chromatin / DNA repair / DNA-templated transcription / DNA damage response / DNA binding ...HU-DNA complex / DnaA-HU complex / bacterial nucleoid DNA packaging / chromosome condensation / DNA replication initiation / structural constituent of chromatin / DNA repair / DNA-templated transcription / DNA damage response / DNA binding / identical protein binding / membrane / cytosol Similarity search - Function |
| Biological species |   |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDGC3 SASDGC3 |
|---|
-Related structure data
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: DNA-binding protein HU-alpha, E34K mutant bound to 80 bp DNA (pH 5.5) Entity id: 1622 / 1623 / 1632 |
|---|---|
| Buffer | Name: 10 mM Bis-Tris, 50 mM NaCl / pH: 5.5 |
| Entity #1622 | Type: DNA / Description: 80bp_DNA Forward / Formula weight: 24.849 / Num. of mol.: 1 / Source: Escherichia coli Sequence: AATGAGGTAA CAACGAAAGC AGATGATAGC TGCTTATCAA TTTGTTGCAA ACAAGTAGCC GCGCCCAATG AGGTAACAAT |
| Entity #1623 | Type: DNA / Description: 80bp_DNA Reverse / Formula weight: 24.612 / Num. of mol.: 1 / Source: Escherichia coli K-12 Sequence: TTACTCCATT GTTGCTTTCG TCTACTATCG ACGAATAGTT AAACAACGTT TGTTCATCGG CGCGGGTTAC TCCATTGTTA |
| Entity #1632 | Type: protein / Description: DNA-binding protein HU-alpha, E34K / Formula weight: 9.534 / Num. of mol.: 2 / Source: Escherichia coli K-12 / References: UniProt: P0ACF0 Sequence: MNKTQLIDVI AEKAELSKTQ AKAALESTLA AITKSLKEGD AVQLVGFGTF KVNHRAERTG RNPQTGKEIK IAAANVPAFV SGKALKDAVK |
-Experimental information
| Beam | Instrument name: Advanced Light Source (ALS) 12.3.1 (SIBYLS) City: Berkeley, CA / 国: USA  / Type of source: X-ray synchrotron / Wavelength: 0.103 Å / Dist. spec. to detc.: 1.5 mm / Type of source: X-ray synchrotron / Wavelength: 0.103 Å / Dist. spec. to detc.: 1.5 mm | ||||||
|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus3 X 2M / Pixsize x: 172 mm | ||||||
| Scan | Measurement date: Nov 2, 2018 / Storage temperature: 10 °C / Cell temperature: 10 °C / Exposure time: 3 sec. / Number of frames: 300 / Unit: 1/A /
| ||||||
| Result | Experimental MW: 69 kDa / Rg from PR: 2 nm / Type of curve: single_conc Comments: his curve is part of a pH series. Increasing the pH removes aggregation presumably due to protein surface charge changes. The SAXS data for this entry is measured from aggregated material ...Comments: his curve is part of a pH series. Increasing the pH removes aggregation presumably due to protein surface charge changes. The SAXS data for this entry is measured from aggregated material and therefore the Rg and I(0) parameters do not apply. |
 Movie
Movie Controller
Controller


