[English] 日本語
 Yorodumi
Yorodumi- SASDF48: Myotillin immuniglobulin domains Ig1Ig2 (250-498) (Myotillin Ig1I... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 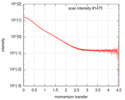 |
|---|---|
 Sample Sample | Myotillin immuniglobulin domains Ig1Ig2 (250-498)
|
| Function / homology |  Function and homology information Function and homology informationaxon guidance receptor activity / structural constituent of muscle / homophilic cell-cell adhesion / alpha-actinin binding / muscle contraction / synapse organization / sarcolemma / Z disc / actin cytoskeleton / actin binding ...axon guidance receptor activity / structural constituent of muscle / homophilic cell-cell adhesion / alpha-actinin binding / muscle contraction / synapse organization / sarcolemma / Z disc / actin cytoskeleton / actin binding / axon / neuronal cell body / plasma membrane Similarity search - Function |
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDF48 SASDF48 |
|---|
-Related structure data
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
|---|
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: Myotillin immuniglobulin domains Ig1Ig2 (250-498) / Specimen concentration: 15.2 mg/ml |
|---|---|
| Buffer | Name: 20 mM Na+-HEPES, 150 mM, NaCl, 5 % v/v glycerol, 1 mM DTT pH: 7.4 |
| Entity #1750 | Name: Ig1Ig2 (250-498) / Type: protein / Description: Myotillin Ig1Ig2 (250-498) / Formula weight: 28.31 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: Q9UBF9 Sequence: GPAMPRFIQV PENMSIDEGR FCRMDFKVSG LPAPDVSWYL NGRTVQSDDL HKMIVSEKGL HSLIFEVVRA SDAGAYACVA KNRAGEATFT VQLDVLAKEH KRAPMFIYKP QSKKVLEGDS VKLECQISAI PPPKLFWKRN NEMVQFNTDR ISLYQDNTGR VTLLIKDVNK ...Sequence: GPAMPRFIQV PENMSIDEGR FCRMDFKVSG LPAPDVSWYL NGRTVQSDDL HKMIVSEKGL HSLIFEVVRA SDAGAYACVA KNRAGEATFT VQLDVLAKEH KRAPMFIYKP QSKKVLEGDS VKLECQISAI PPPKLFWKRN NEMVQFNTDR ISLYQDNTGR VTLLIKDVNK KDAGWYTVSA VNEAGVTTCN TRLDVTARPN QTLPAPKQLR VRPTFSKYLA LNGKGLNVKQ AFNPEGEFQR LAAQSGLYES EEL |
-Experimental information
| Beam | Instrument name: ESRF BM29 / City: Grenoble / 国: France  / Type of source: X-ray synchrotron / Wavelength: 0.099 Å / Dist. spec. to detc.: 2.867 mm / Type of source: X-ray synchrotron / Wavelength: 0.099 Å / Dist. spec. to detc.: 2.867 mm | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 1M / Type: Dectris / Pixsize x: 172 mm | |||||||||||||||||||||||||||||||||
| Scan |
| |||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller




