[English] 日本語
 Yorodumi
Yorodumi- SASDAM6: Naringenin-bound chalcone isomerase (Chalcone isomerase with Nari... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 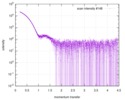 |
|---|---|
 Sample Sample | Naringenin-bound chalcone isomerase
|
| Function / homology | Chalcone isomerase, N-terminal / Chalcone isomerase N-terminal domain / chalcone isomerase / chalcone isomerase activity / Chalcone isomerase Function and homology information Function and homology information |
| Biological species |  Eubacterium ramulus (bacteria) Eubacterium ramulus (bacteria) |
 Citation Citation |  Journal: Acta Crystallogr D Biol Crystallogr / Year: 2015 Journal: Acta Crystallogr D Biol Crystallogr / Year: 2015Title: Structure and catalytic mechanism of the evolutionarily unique bacterial chalcone isomerase. Authors: Maren Thomsen / Anne Tuukkanen / Jonathan Dickerhoff / Gottfried J Palm / Hanna Kratzat / Dmitri I Svergun / Klaus Weisz / Uwe T Bornscheuer / Winfried Hinrichs /  Abstract: Flavonoids represent a large class of secondary metabolites produced by plants. These polyphenolic compounds are well known for their antioxidative abilities, are antimicrobial phytoalexins ...Flavonoids represent a large class of secondary metabolites produced by plants. These polyphenolic compounds are well known for their antioxidative abilities, are antimicrobial phytoalexins responsible for flower pigmentation to attract pollinators and, in addition to other properties, are also specific bacterial regulators governing the expression of Rhizobium genes involved in root nodulation (Firmin et al., 1986). The bacterial chalcone isomerase (CHI) from Eubacterium ramulus catalyses the first step in a flavanone-degradation pathway by ring opening of (2S)-naringenin to form naringenin chalcone. The structural biology and enzymology of plant CHIs have been well documented, whereas the existence of bacterial CHIs has only recently been elucidated. This first determination of the structure of a bacterial CHI provides detailed structural insights into the key step of the flavonoid-degradation pathway. The active site could be confirmed by co-crystallization with the substrate (2S)-naringenin. The stereochemistry of the proposed mechanism of the isomerase reaction was verified by specific (1)H/(2)H isotope exchange observed by (1)H NMR experiments and was further supported by mutagenesis studies. The active site is shielded by a flexible lid, the varying structure of which could be modelled in different states of the catalytic cycle using small-angle X-ray scattering data together with the crystallographic structures. Comparison of bacterial CHI with the plant enzyme from Medicago sativa reveals that they have unrelated folds, suggesting that the enzyme activity evolved convergently from different ancestor proteins. Despite the lack of any functional relationship, the tertiary structure of the bacterial CHI shows similarities to the ferredoxin-like fold of a chlorite dismutase and the stress-related protein SP1. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Models
- Sample
Sample
 Sample Sample | Name: Naringenin-bound chalcone isomerase / Contrast: 3.047 / Specimen concentration: 0.77-2.60 |
|---|---|
| Buffer | Name: 50 mM sodium phosphate / pH: 6.8 |
| Entity #511 | Name: naringenin-CHI / Type: protein / Description: Chalcone isomerase with Naringenin / Formula weight: 32.4 / Num. of mol.: 6 / Source: Eubacterium ramulus / References: UniProt: V9P0A9 Sequence: MADFKFEPMR SLIYVDCVSE DYRPKLQRWI YKVHIPDSIS QFEPYVTKYA FYPSFPIPPQ GDRFGYARMQ LTEHHWLVSD LDPRLEIKAI AETFPMDVLV WQGQIPAAAH TDAQIDSDGD AGNAARKSNN AEGNPFIFAF LPMWWEKDLK GKGRTIEDGA NYRFNMTIGF ...Sequence: MADFKFEPMR SLIYVDCVSE DYRPKLQRWI YKVHIPDSIS QFEPYVTKYA FYPSFPIPPQ GDRFGYARMQ LTEHHWLVSD LDPRLEIKAI AETFPMDVLV WQGQIPAAAH TDAQIDSDGD AGNAARKSNN AEGNPFIFAF LPMWWEKDLK GKGRTIEDGA NYRFNMTIGF PEGVDKAEGE KWLFEKVVPI LQAAPECTRV LASAVKKDIN GCVMDWVLEI WFENQSGWYK VMVDDMKALE KPSWAQQDAF PFLKPYHNVC SAAVADYTPS NNLANYRGYI TMR |
-Experimental information
| Beam | Instrument name: PETRA III P12 / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | ||||||||||||||||||||||||||||||
| Scan |
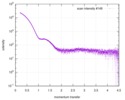
| ||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||
| Result | Comments: Concentrations used: 13.5 µM CHI, 1 mM naringenin.
|
 Movie
Movie Controller
Controller


 SASDAM6
SASDAM6




