[English] 日本語
 Yorodumi
Yorodumi- PDB-8q84: Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8q84 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outer kinetochore Dam1 protomer dimer Ndc80-Nuf2 coiled-coil complex | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | CELL CYCLE / Kinetochore / microtubule / error correction / chromosome segregation | |||||||||
| Function / homology |  Function and homology information Function and homology informationmitotic spindle polar microtubule / Ndc80 complex / DASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / kinetochore organization / positive regulation of attachment of spindle microtubules to kinetochore / meiotic chromosome segregation / mitotic spindle pole body / mitotic spindle midzone ...mitotic spindle polar microtubule / Ndc80 complex / DASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / kinetochore organization / positive regulation of attachment of spindle microtubules to kinetochore / meiotic chromosome segregation / mitotic spindle pole body / mitotic spindle midzone / attachment of spindle microtubules to kinetochore / condensed chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / microtubule plus-end binding / spindle pole body / protein localization to kinetochore / positive regulation of microtubule polymerization / mitotic spindle organization / chromosome segregation / spindle microtubule / mitotic spindle / kinetochore / spindle pole / microtubule / cell division / protein-containing complex binding / identical protein binding / nucleus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Muir, K.W. / Barford, D. | |||||||||
| Funding support |  United Kingdom, 2items United Kingdom, 2items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structural mechanism of outer kinetochore Dam1-Ndc80 complex assembly on microtubules. Authors: Kyle W Muir / Christopher Batters / Tom Dendooven / Jing Yang / Ziguo Zhang / Alister Burt / David Barford /  Abstract: Kinetochores couple chromosomes to the mitotic spindle to segregate the genome during cell division. An error correction mechanism drives the turnover of kinetochore-microtubule attachments until ...Kinetochores couple chromosomes to the mitotic spindle to segregate the genome during cell division. An error correction mechanism drives the turnover of kinetochore-microtubule attachments until biorientation is achieved. The structural basis for how kinetochore-mediated chromosome segregation is accomplished and regulated remains an outstanding question. In this work, we describe the cryo-electron microscopy structure of the budding yeast outer kinetochore Ndc80 and Dam1 ring complexes assembled onto microtubules. Complex assembly occurs through multiple interfaces, and a staple within Dam1 aids ring assembly. Perturbation of key interfaces suppresses yeast viability. Force-rupture assays indicated that this is a consequence of impaired kinetochore-microtubule attachment. The presence of error correction phosphorylation sites at Ndc80-Dam1 ring complex interfaces and the Dam1 staple explains how kinetochore-microtubule attachments are destabilized and reset. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8q84.cif.gz 8q84.cif.gz | 608.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8q84.ent.gz pdb8q84.ent.gz | Display |  PDB format PDB format | |
| PDBx/mmJSON format |  8q84.json.gz 8q84.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  8q84_validation.pdf.gz 8q84_validation.pdf.gz | 1.3 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  8q84_full_validation.pdf.gz 8q84_full_validation.pdf.gz | 1.3 MB | Display | |
| Data in XML |  8q84_validation.xml.gz 8q84_validation.xml.gz | 72 KB | Display | |
| Data in CIF |  8q84_validation.cif.gz 8q84_validation.cif.gz | 122 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q8/8q84 https://data.pdbj.org/pub/pdb/validation_reports/q8/8q84 ftp://data.pdbj.org/pub/pdb/validation_reports/q8/8q84 ftp://data.pdbj.org/pub/pdb/validation_reports/q8/8q84 | HTTPS FTP |
-Related structure data
| Related structure data | 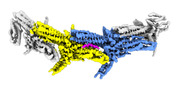 18246MC  8q85C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Kinetochore protein ... , 2 types, 4 molecules AFBG
| #1: Protein | Mass: 80609.375 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: NDC80 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P40460 Trichoplusia ni (cabbage looper) / References: UniProt: P40460#2: Protein | Mass: 53025.949 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: NUF2 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P33895 Trichoplusia ni (cabbage looper) / References: UniProt: P33895 |
|---|
-DASH complex subunit ... , 10 types, 21 molecules IUeJVKWLXMYNZOaPbQcRd
| #3: Protein | Mass: 38477.871 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DAM1 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P53267 Trichoplusia ni (cabbage looper) / References: UniProt: P53267#4: Protein | Mass: 27515.008 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DUO1 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P53168 Trichoplusia ni (cabbage looper) / References: UniProt: P53168#5: Protein | Mass: 15084.501 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DAD2 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P36162 Trichoplusia ni (cabbage looper) / References: UniProt: P36162#6: Protein | Mass: 10522.616 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DAD1 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: Q12248 Trichoplusia ni (cabbage looper) / References: UniProt: Q12248#7: Protein | Mass: 8164.255 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DAD4 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P69851 Trichoplusia ni (cabbage looper) / References: UniProt: P69851#8: Protein | Mass: 10862.234 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: DAD3 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P69850 Trichoplusia ni (cabbage looper) / References: UniProt: P69850#9: Protein | Mass: 34131.812 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: SPC34 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P36131 Trichoplusia ni (cabbage looper) / References: UniProt: P36131#10: Protein | Mass: 32106.631 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: ASK1 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P35734 Trichoplusia ni (cabbage looper) / References: UniProt: P35734#11: Protein | Mass: 8097.205 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: HSK3 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: P69852 Trichoplusia ni (cabbage looper) / References: UniProt: P69852#12: Protein | Mass: 18935.559 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: SPC19 / Production host:  Trichoplusia ni (cabbage looper) / References: UniProt: Q03954 Trichoplusia ni (cabbage looper) / References: UniProt: Q03954 |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Outer kinetochore Dam1 protomer dimer with staple and Ndc80-Nuf2 coiled-coils Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 673772.36 / Experimental value: NO |
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Buffer solution | pH: 6.8 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: TFS KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 3000 nm / Nominal defocus min: 1200 nm |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) |
- Processing
Processing
| EM software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 248732 / Symmetry type: POINT | ||||||||||||||||||||||||||||
| Atomic model building | B value: 540.98 / Protocol: OTHER / Space: REAL Details: Initial rigid body fitting was performed in chimera, with manual correction in coot and real-space refinement in PHENIX | ||||||||||||||||||||||||||||
| Atomic model building | Source name: AlphaFold / Type: in silico model | ||||||||||||||||||||||||||||
| Refinement | Cross valid method: NONE Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2 | ||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 540.98 Å2 | ||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj

